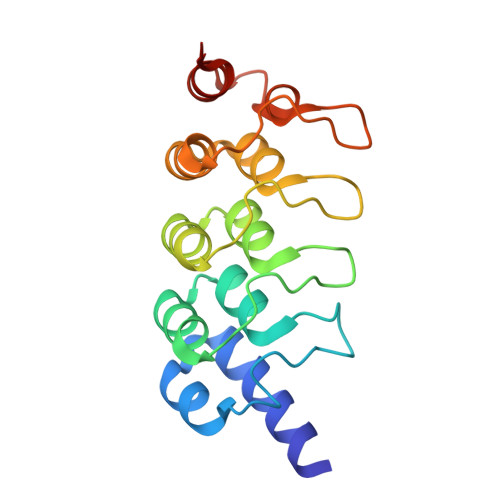
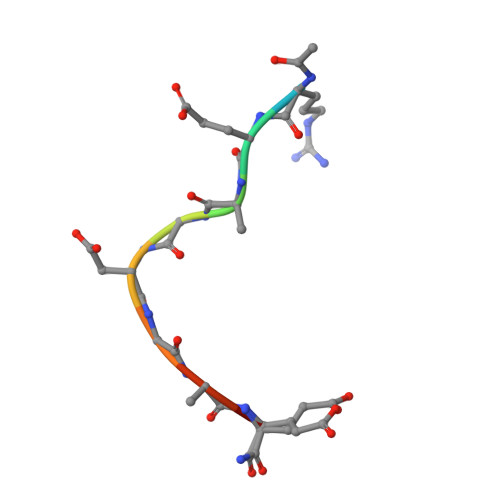
Macrocyclized Extended Peptides: Inhibiting the Substrate-Recognition Domain of Tankyrase.
Xu, W., Lau, Y.H., Fischer, G., Tan, Y.S., Chattopadhyay, A., de la Roche, M., Hyvonen, M., Verma, C., Spring, D.R., Itzhaki, L.S.(2017) J Am Chem Soc 139: 2245-2256
- PubMed: 28084734 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.6b10234
- Primary Citation Related Structures:
5BXO, 5BXU - PubMed Abstract:
We report a double-click macrocyclization approach for the design of constrained peptide inhibitors having non-helical or extended conformations. Our targets are the tankyrase proteins (TNKS), poly(ADP-ribose) polymerases (PARP) that regulate Wnt signaling by targeting Axin for degradation. TNKS are deregulated in many different cancer types, and inhibition of TNKS therefore represents an attractive therapeutic strategy. However, clinical development of TNKS-specific PARP catalytic inhibitors is challenging due to off-target effects and cellular toxicity. We instead targeted the substrate-recognition domain of TNKS, as it is unique among PARP family members. We employed a two-component strategy, allowing peptide and linker to be separately engineered and then assembled in a combinatorial fashion via click chemistry. Using the consensus substrate-peptide sequence as a starting point, we optimized the length and rigidity of the linker and its position along the peptide. Optimization was further guided by high-resolution crystal structures of two of the macrocyclized peptides in complex with TNKS. This approach led to macrocyclized peptides with submicromolar affinities for TNKS and high proteolytic stability that are able to disrupt the interaction between TNKS and Axin substrate and to inhibit Wnt signaling in a dose-dependent manner. The peptides therefore represent a promising starting point for a new class of substrate-competitive inhibitors of TNKS with potential for suppressing Wnt signaling in cancer. Moreover, by demonstrating the application of the double-click macrocyclization approach to non-helical, extended, or irregularly structured peptides, we greatly extend its potential and scope, especially given the frequency with which such motifs mediate protein-protein interactions.
- Department of Pharmacology, University of Cambridge , Tennis Court Road, Cambridge CB2 1PD, United Kingdom.
Organizational Affiliation: