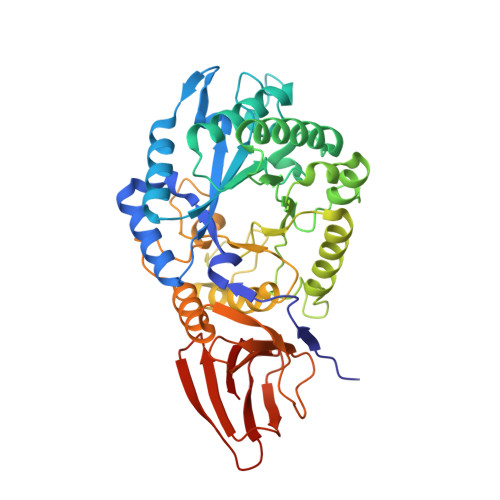
Characterization of the Pseudomonas aeruginosa Glycoside Hydrolase PslG Reveals That Its Levels Are Critical for Psl Polysaccharide Biosynthesis and Biofilm Formation.
Baker, P., Whitfield, G.B., Hill, P.J., Little, D.J., Pestrak, M.J., Robinson, H., Wozniak, D.J., Howell, P.L.(2015) J Biological Chem 290: 28374-28387
- PubMed: 26424791 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.674929
- Primary Citation Related Structures:
5BX9, 5BXA - PubMed Abstract:
A key component of colonization, biofilm formation, and protection of the opportunistic human pathogen Pseudomonas aeruginosa is the biosynthesis of the exopolysaccharide Psl. Composed of a pentameric repeating unit of mannose, glucose, and rhamnose, the biosynthesis of Psl is proposed to occur via a Wzx/Wzy-dependent mechanism. Previous genetic studies have shown that the putative glycoside hydrolase PslG is essential for Psl biosynthesis. To understand the function of this protein, the apo-structure of the periplasmic domain of PslG (PslG(31-442)) and its complex with mannose were determined to 2.0 and 1.9 Å resolution, respectively. Despite a domain architecture and positioning of catalytic residues similar to those of other family 39 glycoside hydrolases, PslG(31-442) exhibits a unique 32-Å-long active site groove that is distinct from other structurally characterized family members. PslG formed a complex with two mannose monosaccharides in this groove, consistent with binding data obtained from intrinsic tryptophan fluorescence. PslG was able to catalyze the hydrolysis of surface-associated Psl, and this activity was abolished in a E165Q/E276Q double catalytic variant. Surprisingly, P. aeruginosa variants with these chromosomal mutations as well as a pslG deletion mutant were still capable of forming Psl biofilms. However, overexpression of PslG in a pslG deletion background impaired biofilm formation and resulted in less surface-associated Psl, suggesting that regulation of this enzyme is important during polysaccharide biosynthesis.
- Program in Molecular Structure and Function, Research Institute, The Hospital for Sick Children, Toronto, Ontario M5G 1X8, Canada.
Organizational Affiliation: