Synthesis and Pharmacology of Mono-, Di-, and Trialkyl-Substituted 7-Chloro-3,4-dihydro-2H-1,2,4-benzothiadiazine 1,1-Dioxides Combined with X-ray Structure Analysis to Understand the Unexpected Structure-Activity Relationship at AMPA Receptors.
Larsen, A.P., Francotte, P., Frydenvang, K., Tapken, D., Goffin, E., Fraikin, P., Caignard, D.H., Lestage, P., Danober, L., Pirotte, B., Kastrup, J.S.(2016) ACS Chem Neurosci 7: 378-390
- PubMed: 26771108 Search on PubMed
- DOI: https://doi.org/10.1021/acschemneuro.5b00318
- Primary Citation Related Structures:
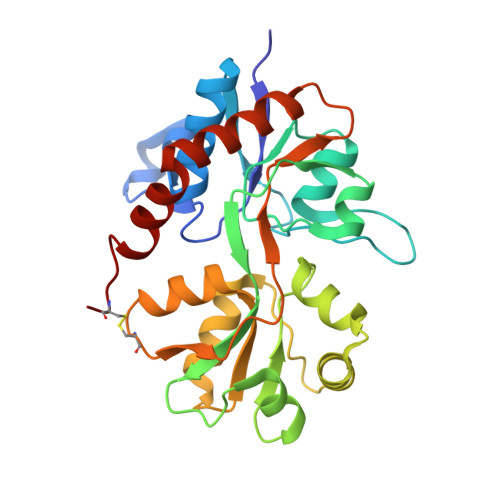
5BUU - PubMed Abstract:
Positive allosteric modulators of 2-amino-3-(3-hydroxy-5-methylisoxazol-4-yl)propionic acid (AMPA)-type ionotropic glutamate receptors are promising compounds for treatment of neurological disorders, for example, Alzheimer's disease. Here, we report synthesis and pharmacological evaluation of a series of mono-, di-, or trialkyl-substituted 7-chloro-3,4-dihydro-2H-1,2,4-benzothiadiazine 1,1-dioxides, comprising in total 16 new modulators. The trisubstituted compounds 7b, 7d, and 7e revealed potent activity (EC2× = 2.7-4.3 μM; concentration of compound responsible for a 2-fold increase of the AMPA mediated response) as AMPA receptor potentiators in an in vitro cellular fluorescence assay (FLIPR). The 4-cyclopropyl compound 7f was found to be considerably less potent (EC2× = 60 μM), in contrast to previously described 4-monoalkyl-substituted benzothiadiazine dioxides for which the cyclopropyl group constitutes the best choice of substituent. 7b was subjected to X-ray structural analysis in complex with the GluA2 ligand-binding domain. We propose an explanation of the unexpected structure-activity relationship of this new series of mono-, di-, and trialkyl-substituted 1,2,4-benzothiadiazine 1,1-dioxide compounds. The methyl substituent in the 3-position directs the binding mode of the 1,2,4-benzothiadiazine 1,1-dioxide (BTD) scaffold. When a methyl substituent is present in the 3-position of the BTD, additional methyl substituents in both the 2- and 4-positions increase potency, whereas introduction of a 4-cyclopropyl group does not enhance potency of 2,3,4-alkyl-substituted BTDs. A hydrogen bond donor in the 2-position of the BTD is not necessary for modulator potency.
- Department of Drug Design and Pharmacology, Faculty of Health and Medical Sciences, University of Copenhagen , Universitetsparken, 2, DK-2100 Copenhagen, Denmark.
Organizational Affiliation: