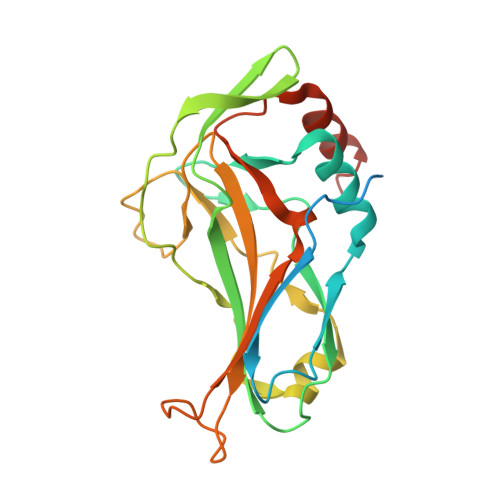
Intermolecular Interactions of Cardiac Transcription Factors NKX2.5 and TBX5.
Pradhan, L., Gopal, S., Li, S., Ashur, S., Suryanarayanan, S., Kasahara, H., Nam, H.J.(2016) Biochemistry 55: 1702-1710
- PubMed: 26926761 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.6b00171
- Primary Citation Related Structures:
4S0H, 5BQD - PubMed Abstract:
Heart development in mammalian systems is controlled by combinatorial interactions of master cardiac transcription factors such as TBX5 and NKX2.5. They bind to promoters/enhancers of downstream targets as homo- or heteromultimeric complexes. They physically interact and synergistically regulate their target genes. To elucidate the molecular basis of the intermolecular interactions, a heterodimer and a homodimer of NKX2.5 and TBX5 were studied using X-ray crystallography. Here we report a crystal structure of human NKX2.5 and TBX5 DNA binding domains in a complex with a 19 bp target DNA and a crystal structure of TBX5 homodimer. The ternary complex structure of NKX2.5 and TBX5 with the target DNA shows physical interactions between the two proteins through Lys158 (NKX2.5), Asp140 (TBX5), and Pro142 (TBX5), residues that are highly conserved in TBX and NKX families across species. Extensive homodimeric interactions were observed between the TBX5 proteins in both crystal structures. In particular, in the crystal structure of TBX5 protein that includes the N-terminal and DNA binding domains, intermolecular interactions were mediated by the N-terminal domain of the protein. The N-terminal domain of TBX5 was predicted to be "intrinsically unstructured", and in one of the two molecules in an asymmetric unit, the N-terminal domain assumes a β-strand conformation bridging two β-sheets from the two molecules. The structures reported here may represent general mechanisms for combinatorial interactions among transcription factors regulating developmental processes.
- Department of Bioengineering, University of Texas at Dallas , Richardson, Texas 75080, United States.
Organizational Affiliation: