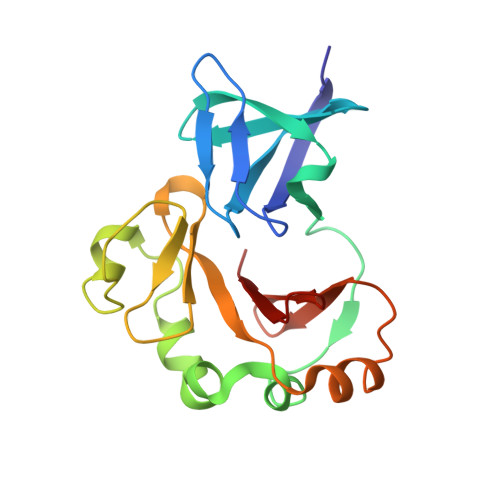
SH3-Like Motif-Containing C-terminal Domain of Staphylococcal Teichoic Acid Transporter Suggests Possible Function.
Ko, T.P., Tseng, S.T., Lai, S.J., Chen, S.C., Guan, H.H., Yang, C.S., Chen, C.J., Chen, Y.(2016) Proteins
- PubMed: 27213893 Search on PubMed
- DOI: https://doi.org/10.1002/prot.25074
- Primary Citation Related Structures:
5BND - PubMed Abstract:
The negatively charged bacterial polysaccharides-wall teichoic acids (WTAs) are synthesized intracellularly and exported by a two-component transporter, TagGH, comprising a transmembrane subunit TagG and an ATPase subunit TagH. We determined the crystal structure of the C-terminal domain of TagH (TagH-C) to investigate its function. The structure shows an N-terminal SH3-like subdomain wrapped by a C-terminal subdomain with an anti-parallel β-sheet and an outer shell of α-helices. A stretch of positively charged surface across the subdomain interface is flanked by two negatively charged regions, suggesting a potential binding site for negatively charged polymers, such as WTAs or acidic peptide chains. Proteins 2016; 84:1328-1332. © 2016 Wiley Periodicals, Inc.
- Institute of Biological Chemistry, Academia Sinica, Taipei, Taiwan.
Organizational Affiliation: