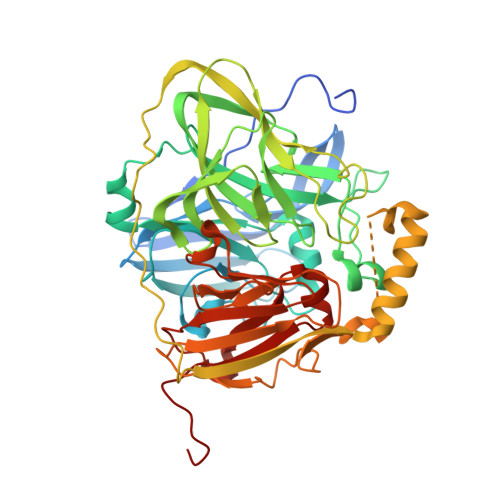
Biochemical, spectroscopic and X-ray structural analysis of deuterated multicopper oxidase CueO prepared from a new expression construct for neutron crystallography
Akter, M., Inoue, C., Komori, H., Matsuda, N., Sakurai, T., Kataoka, K., Higuchi, Y., Shibata, N.(2016) Acta Crystallogr F Struct Biol Commun 72: 788-794
- PubMed: 27710945 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X1601400X
- Primary Citation Related Structures:
5B7E, 5B7F, 5B7M - PubMed Abstract:
Multicopper oxidases oxidize various phenolic and nonphenolic compounds by using molecular oxygen as an electron acceptor to produce water. A multicopper oxidase protein, CueO, from Escherichia coli is involved in copper homeostasis in the bacterial cell. Although X-ray crystallographic studies have been conducted, the reduction mechanism of oxygen and the proton-transfer pathway remain unclear owing to the difficulty in identifying H atoms from X-ray diffraction data alone. To elucidate the reaction mechanism using neutron crystallography, a preparation system for obtaining large, high-quality single crystals of deuterated CueO was developed. Tiny crystals were obtained from the deuterated CueO initially prepared from the original construct. The X-ray crystal structure of the deuterated CueO showed that the protein contained an incompletely truncated signal sequence at the N-terminus, which resulted in the heterogeneity of the protein sample for crystallization. Here, a new CueO expression system that had an HRV3C cleavage site just after the signal sequence was constructed. Deuterated CueO from the new construct was expressed in cells cultured in deuterated algae-extract medium and the signal sequence was completely eliminated by HRV3C protease. The deuteration level of the purified protein was estimated by MALDI-TOF mass spectrometry to be at least 83.2% compared with nondeuterated protein. Nondeuterated CueO crystallized in space group P2 1 , with unit-cell parameters a = 49.51, b = 88.79, c = 53.95 Å, β = 94.24°, and deuterated CueO crystallized in space group P2 1 2 1 2 1 , with unit-cell parameters a = 49.91, b = 106.92, c = 262.89 Å. The crystallographic parameters for the crystals of the new construct were different from those previously reported for nondeuterated crystals. The nondeuterated and deuterated CueO from the new construct had similar UV-Vis spectra, enzymatic activities and overall structure and geometry of the ligands of the Cu atoms in the active site to those of previously reported CueO structures. These results indicate that the CueO protein prepared using the new construct is suitable for further neutron diffraction studies.
- Department of Life Science, Graduate School of Life Science, University of Hyogo, 3-2-1 Koto, Kamigori, Ako-gun, Hyogo 678-1297, Japan.
Organizational Affiliation: