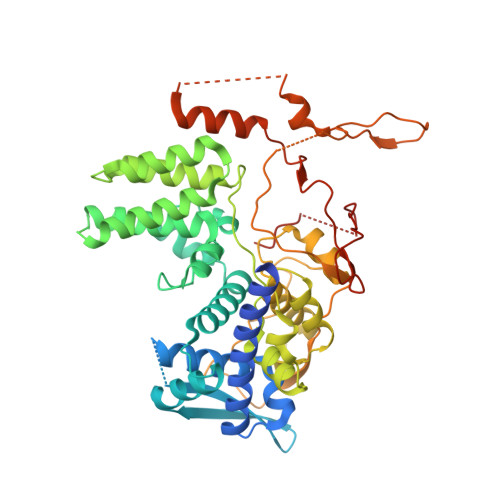
Structural Characterization of H1N1 Nucleoprotein-Nucleozin Binding Sites
Pang, B., Cheung, N.N., Zhang, W.Z., Dai, J., Kao, R.Y., Zhang, H.M., Hao, Q.(2016) Sci Rep 6: 29684-29684
- PubMed: 27404920 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep29684
- Primary Citation Related Structures:
5B7B - PubMed Abstract:
Influenza viruses are among the most common pathogens that threaten the health of humans and animals worldwide. Various anti-viral therapeutic agents are currently used for treatment and prophylaxis of influenza virus, but the targets of these drugs are easily mutated and result in resistance. Therefore, medications that have broad spectrum coverage are urgently needed to combat with the disease. Since nucleoprotein is regarded as a druggable target due to its conserved sequence and important functions during influenza virus life cycle, numerous studies are focused on this protein in attempts to develop broad-spectrum anti-influenza therapeutics. Recently, a novel small molecule compound, nucleozin, was found to induce large aggregates of nucleoprotein, which in turn caused cessation of virus replication. However, the aggregation-inducing mechanism of nucleozin has not been unveiled. Here we report the crystal structure of nucleoprotein-nucleozin complex at 3 Å resolution, which shows the binding sites of nucleozin at nucleoprotein for the first time. The complex structure reveals how nucleoprotein and nucleozin interact with each other and hence result in nucleoprotein aggregates. The structural information is envisaged to help accelerate the development of anti-influenza therapeutic agents.
- School of Biomedical Sciences, The University of Hong Kong, Hong Kong SAR, China.
Organizational Affiliation: