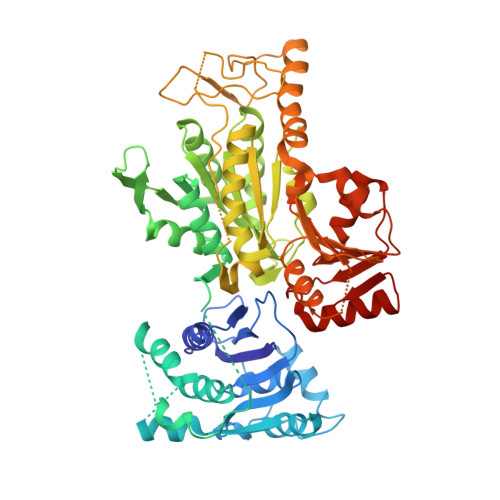
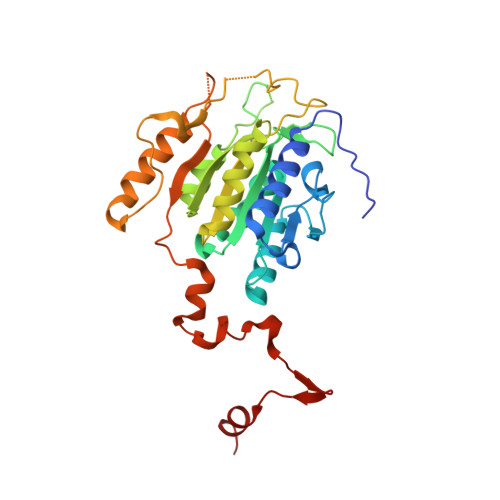
Crystal structures of archaeal 2-oxoacid:ferredoxin oxidoreductases from Sulfolobus tokodaii
Yan, Z., Maruyama, A., Arakawa, T., Fushinobu, S., Wakagi, T.(2016) Sci Rep 6: 33061-33061
- PubMed: 27619895 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep33061
- Primary Citation Related Structures:
5B46, 5B47, 5B48 - PubMed Abstract:
As the first three-dimensional structure of the two-subunit type 2-oxoacid:ferredoxin oxidoreductases (OFOR) from archaea, we solved the crystal structures of STK_23000/STK_22980 (StOFOR1) and STK_24350/STK_24330 (StOFOR2) from Sulfolobus tokodaii. They showed similar overall structures, consisting of two a- and b-subunit heterodimers containing thiamin pyrophosphate (TPP) cofactor and [4Fe-4S] cluster, but lack an intramolecular ferredoxin domain. Unlike other OFORs, StOFORs can utilize both pyruvate and 2-oxoglutarate, playing a key role in the central metabolism. In the structure of StOFOR2 in unreacted pyruvate complex form, carboxylate group of pyruvate is recognized by Arg344 and Thr257 from the a-subunit, which are conserved in pyruvate:ferredoxin oxidoreductase from Desulfovbrio africanus (DaPFOR). In the structure of StOFOR1 co-crystallized with 2-oxobutyrate, electron density corresponding to a 1-hydroxypropyl group (post-decarboxylation state) was observed at the thiazole ring of TPP. The binding pockets of the StOFORs surrounding the methyl or propyl group of the ligands are wider than that of DaPFOR. Mutational analyses indicated that several residues were responsible for the broad 2-oxoacid specificity of StOFORs. We also constructed a possible complex structural model by placing a Zn(2+)-containing dicluster ferredoxin of S. tokodaii into the large pocket of StOFOR2, providing insight into the electron transfer between the two redox proteins.
- Department of Biotechnology, Graduate School of Agricultural and Life Sciences, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.
Organizational Affiliation: