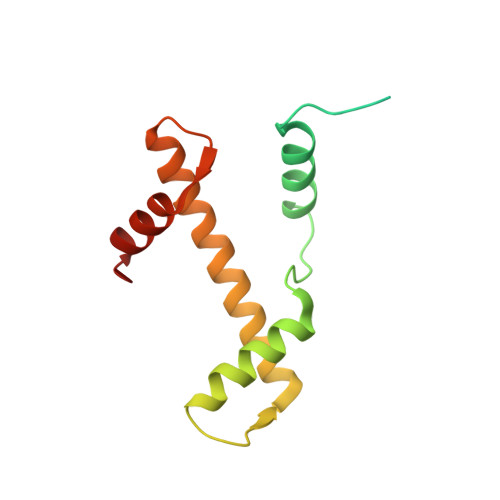
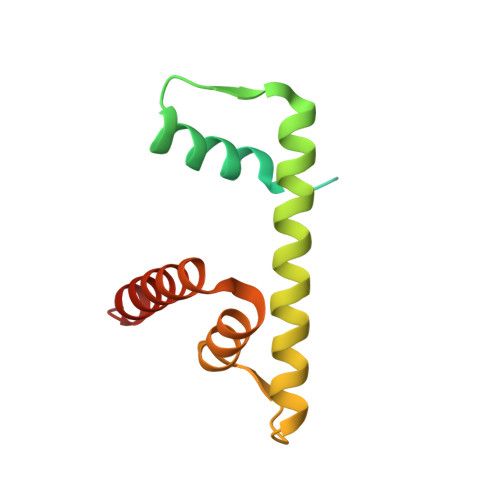
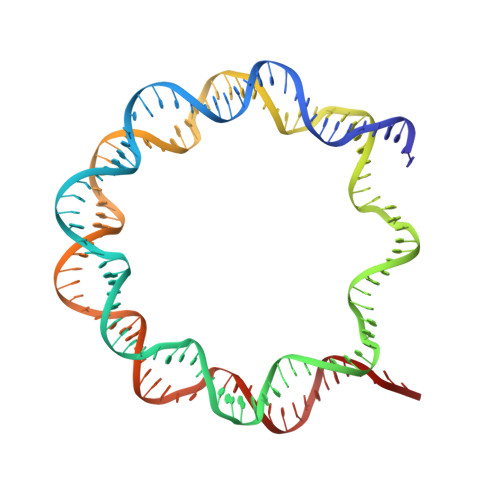
Crystal structures of heterotypic nucleosomes containing histones H2A.Z and H2A.
Horikoshi, N., Arimura, Y., Taguchi, H., Kurumizaka, H.(2016) Open Biol 6
- PubMed: 27358293
- DOI: https://doi.org/10.1098/rsob.160127
- Primary Citation Related Structures:
5B31, 5B32, 5B33 - PubMed Abstract:
H2A.Z is incorporated into nucleosomes located around transcription start sites and functions as an epigenetic regulator for the transcription of certain genes. During transcriptional regulation, the heterotypic H2A.Z/H2A nucleosome containing one each of H2A.Z and H2A is formed. However, previous homotypic H2A.Z nucleosome structures suggested that the L1 loop region of H2A.Z would sterically clash with the corresponding region of canonical H2A in the heterotypic nucleosome. To resolve this issue, we determined the crystal structures of heterotypic H2A.Z/H2A nucleosomes. In the H2A.Z/H2A nucleosome structure, the H2A.Z L1 loop structure was drastically altered without any structural changes of the canonical H2A L1 loop, thus avoiding the steric clash. Unexpectedly, the heterotypic H2A.Z/H2A nucleosome is more stable than the homotypic H2A.Z nucleosome. These data suggested that the flexible character of the H2A.Z L1 loop plays an essential role in forming the stable heterotypic H2A.Z/H2A nucleosome.
- Research Institute for Science and Engineering, Waseda University, 2-2 Wakamatsu-cho, Shinjuku-ku, Tokyo 162-8480, Japan.
Organizational Affiliation: