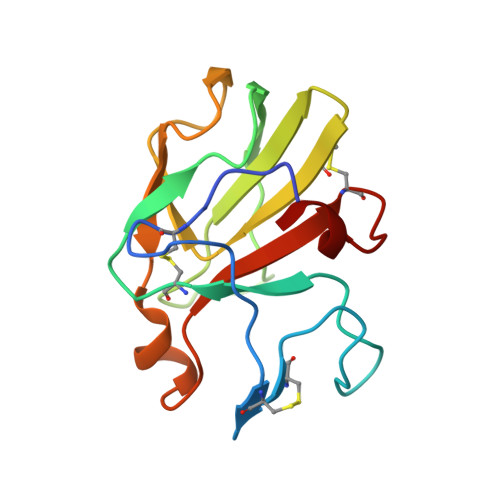
Crystal Structure of Human Leukocyte Cell-derived Chemotaxin 2 (LECT2) Reveals a Mechanistic Basis of Functional Evolution in a Mammalian Protein with an M23 Metalloendopeptidase Fold
Zheng, H., Miyakawa, T., Sawano, Y., Asano, A., Okumura, A., Yamagoe, S., Tanokura, M.(2016) J Biological Chem 291: 17133-17142
- PubMed: 27334921 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M116.720375
- Primary Citation Related Structures:
5B0H - PubMed Abstract:
Human leukocyte cell-derived chemotaxin 2 (LECT2), which is predominantly expressed in the liver, is a multifunctional protein. LECT2 is becoming a potential therapeutic target for several diseases of worldwide concern such as rheumatoid arthritis, hepatocellular carcinoma, and obesity. Here, we present the crystal structure of LECT2, the first mammalian protein whose structure contains an M23 metalloendopeptidase fold. The LECT2 structure adopts a conserved Zn(II) coordination configuration but lacks a proposed catalytic histidine residue, and its potential substrate-binding groove is blocked in the vicinity of the Zn(II)-binding site by an additional intrachain loop at the N terminus. Consistent with these structural features, LECT2 was found to be catalytically inactive as a metalloendopeptidase against various types of peptide sequences, including pentaglycine. In addition, a surface plasmon resonance analysis demonstrated that LECT2 bound to the c-Met receptor with micromolar affinity. These results indicate that LECT2 likely plays its critical roles by acting as a ligand for the corresponding protein receptors rather than as an enzymatically active peptidase. The intrachain loop together with the pseudo-active site groove in LECT2 structure may be specific for interactions between LECT2 and receptors. Our study reveals a mechanistic basis for the functional evolution of a mammalian protein with an M23 metalloendopeptidase fold and potentially broadens the implications for the biological importance of noncatalytic peptidases in the M23 family.
- From the Department of Applied Biological Chemistry, Graduate School of Agricultural and Life Sciences, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.
Organizational Affiliation: