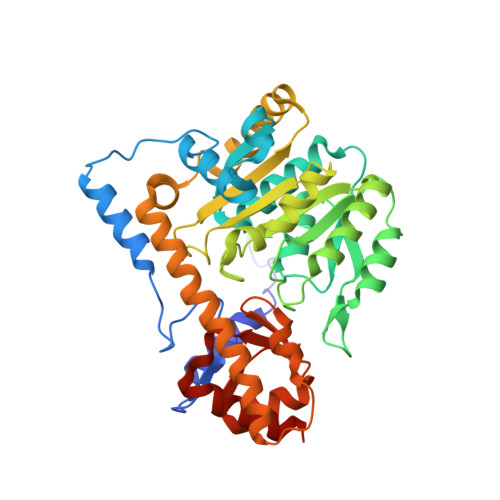
Recombinant expression, purification and crystallographic studies of the mature form of human mitochondrial aspartate aminotransferase
Jiang, X., Wang, J., Chang, H., Zhou, Y.(2016) Biosci Trends 10: 79-84
- PubMed: 26902786 Search on PubMed
- DOI: https://doi.org/10.5582/bst.2015.01150
- Primary Citation Related Structures:
5AX8 - PubMed Abstract:
Mitochondrial aspartate aminotransferase (mAspAT) was recognized as a moonlighting enzyme because it has not only aminotransferase activity but also a high-affinity long-chain fatty acids (LCFA) binding site. This enzyme plays a key role in amino acid metabolism, biosynthesis of kynurenic acid and transport of the LCFA. Therefore, it is important to study the structure-function relationships of human mAspAT protein. In this work, the mature form of human mAspAT was expressed to a high level in Escherichia coli periplasmic space using pET-22b vector, purified by a combination of immobilized metal-affinity chromatography and cation exchange chromatography. Optimal activity of the enzyme occurred at a temperature of 47.5ºC and a pH of 8.5. Crystals of human mAspAT were grown using the hanging-drop vapour diffusion method at 277K with 0.1 M HEPES pH 6.8 and 25%(v/v) Jeffamine(®) ED-2001 pH 6.8. The crystals diffracted to 2.99 Å and belonged to the space group P1 with the unit-cell parameters a =56.7, b = 76.1, c = 94.2 Å, α =78.0, β =85.6, γ = 78.4º. Elucidation of mAspAT structure can provide a molecular basis towards understanding catalysis mechanism and substrate binding site of enzyme.
- School of Life Science and Biotechnology, Dalian University of Technology.
Organizational Affiliation: