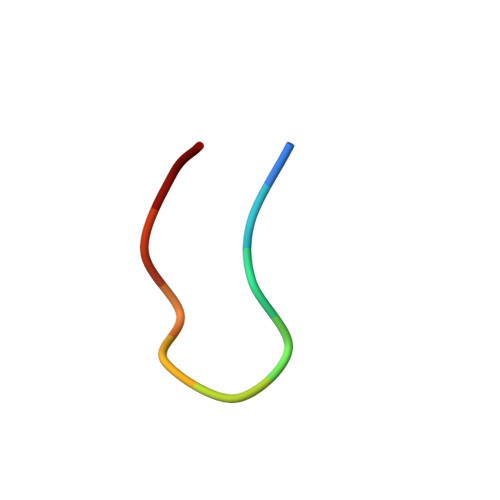Crystal structure of a ten-amino acid protein
Honda, S., Akiba, T., Kato, Y.S., Sawada, Y., Sekijima, M., Ishimura, M., Ooishi, A., Watanabe, H., Odahara, T., Harata, K.(2008) J Am Chem Soc 130: 15327-15331
- PubMed: 18950166 Search on PubMed
- DOI: https://doi.org/10.1021/ja8030533
- Primary Citation Related Structures:
2RVD, 5AWL - PubMed Abstract:
What is the smallest protein? This is actually not such a simple question to answer, because there is no established consensus among scientists as to the definition of a protein. We describe here a designed molecule consisting of only 10 amino acids. Despite its small size, its essential characteristics, revealed by its crystal structure, solution structure, thermal stability, free energy surface, and folding pathway network, are consistent with the properties of natural proteins. The existence of this kind of molecule deepens our understanding of proteins and impels us to define an "ideal protein" without inquiring whether the molecule actually occurs in nature.
- National Institute of Advanced Industrial Science and Technology, Central 6, Tsukuba 305-8566, Japan. s.honda@aist.go.jp
Organizational Affiliation: