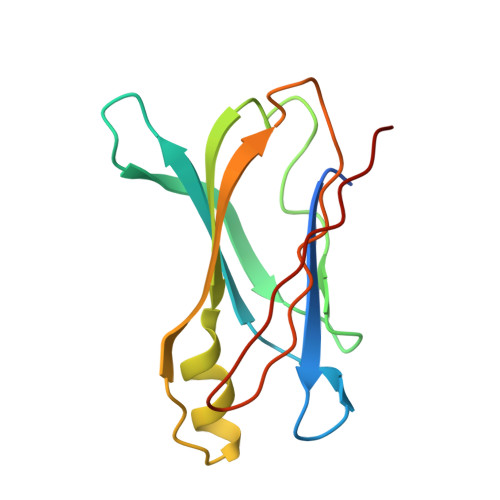
Transthyretin Binding Heterogeneity and Anti-Amyloidogenic Activity of Natural Polyphenols and Their Metabolites
Florio, P., Folli, C., Cianci, M., Del Rio, D., Zanotti, G., Berni, R.(2015) J Biological Chem 290: 29769
- PubMed: 26468275 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.690172
- Primary Citation Related Structures:
5AKS, 5AKT, 5AKV, 5AL0, 5AL8, 5CR1 - PubMed Abstract:
Transthyretin (TTR) is an amyloidogenic protein, the amyloidogenic potential of which is enhanced by a number of specific point mutations. The ability to inhibit TTR fibrillogenesis is known for several classes of compounds, including natural polyphenols, which protect the native state of TTR by specifically interacting with its thyroxine binding sites. Comparative analyses of the interaction and of the ability to protect the TTR native state for polyphenols, both stilbenoids and flavonoids, and some of their main metabolites have been carried out. A main finding of this investigation was the highly preferential binding of resveratrol and thyroxine, both characterized by negative binding cooperativity, to distinct sites in TTR, consistent with the data of x-ray analysis of TTR in complex with both ligands. Although revealing the ability of the two thyroxine binding sites of TTR to discriminate between different ligands, this feature has allowed us to evaluate the interactions of polyphenols with both resveratrol and thyroxine preferential binding sites, by using resveratrol and radiolabeled T4 as probes. Among flavonoids, genistein and apigenin were able to effectively displace resveratrol from its preferential binding site, whereas genistein also showed the ability to interact, albeit weakly, with the preferential thyroxine binding site. Several glucuronidated polyphenol metabolites did not exhibit significant competition for resveratrol and thyroxine preferential binding sites and lacked the ability to stabilize TTR. However, resveratrol-3-O-sulfate was able to significantly protect the protein native state. A rationale for the in vitro properties found for polyphenol metabolites was provided by x-ray analysis of their complexes with TTR.
- From the Department of Life Sciences, University of Parma, 43124 Parma, Italy.
Organizational Affiliation: