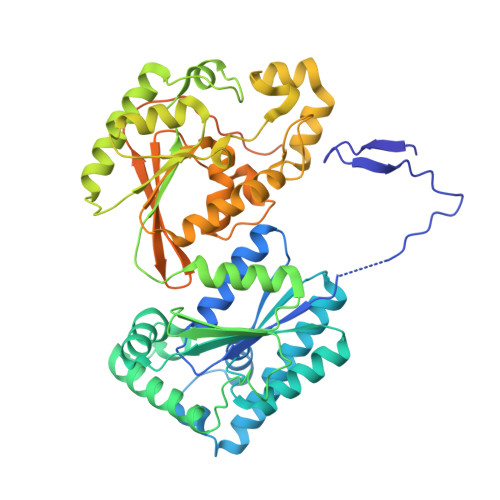
Structure-Based Design of Potent and Selective Inhibitors of the Metabolic Kinase Pfkfb3.
Boyd, S., Brookfield, J.L., Critchlow, S.E., Cumming, I.A., Curtis, N.J., Debreczeni, J., Degorce, S.L., Donald, C., Evans, N.J., Groombridge, S., Hopcroft, P., Jones, N.P., Kettle, J.G., Lamont, S., Lewis, H.J., Macfaull, P., Mcloughlin, S.B., Rigoreau, L.J.M., Smith, J.M., St-Gallay, S., Stock, J.K., Turnbull, A.P., Wheatley, E.R., Winter, J., Wingfield, J.(2015) J Med Chem 58: 3611
- PubMed: 25849762
- DOI: https://doi.org/10.1021/acs.jmedchem.5b00352
- Primary Citation of Related Structures:
5AJV, 5AJW, 5AJX, 5AJY, 5AJZ, 5AK0 - PubMed Abstract:
A weak screening hit with suboptimal physicochemical properties was optimized against PFKFB3 kinase using critical structure-guided insights. The resulting compounds demonstrated high selectivity over related PFKFB isoforms and modulation of the target in a cellular context. A selected example demonstrated exposure in animals following oral dosing. Examples from this series may serve as useful probes to understand the emerging biology of this metabolic target.
- †Oncology Innovative Medicines Unit, AstraZeneca, 35S47 Mereside, Alderley Park, Macclesfield, Cheshire SK10 4TG, United Kingdom.
Organizational Affiliation: