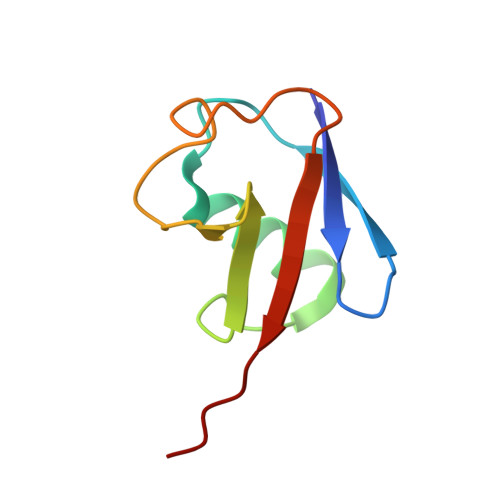
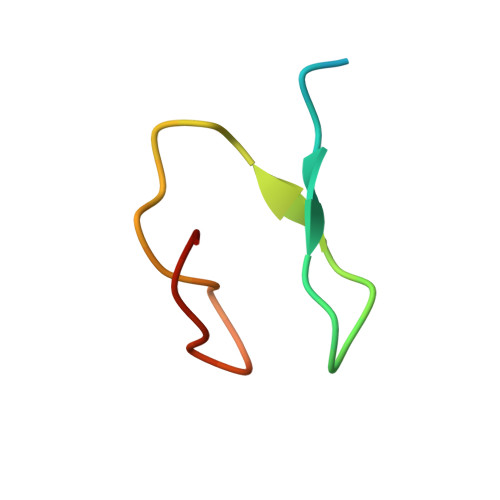
Assembly and Specific Recognition of K29- and K33-Linked Polyubiquitin.
Michel, M.A., Elliott, P.R., Swatek, K.N., Simicek, M., Pruneda, J.N., Wagstaff, J.L., Freund, S.M.V., Komander, D.(2015) Mol Cell 58: 95
- PubMed: 25752577 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2015.01.042
- Primary Citation Related Structures:
5AF4, 5AF5, 5AF6 - PubMed Abstract:
Protein ubiquitination regulates many cellular processes via attachment of structurally and functionally distinct ubiquitin (Ub) chains. Several atypical chain types have remained poorly characterized because the enzymes mediating their assembly and receptors with specific binding properties have been elusive. We found that the human HECT E3 ligases UBE3C and AREL1 assemble K48/K29- and K11/K33-linked Ub chains, respectively, and can be used in combination with DUBs to generate K29- and K33-linked chains for biochemical and structural analyses. Solution studies indicate that both chains adopt open and dynamic conformations. We further show that the N-terminal Npl4-like zinc finger (NZF1) domain of the K29/K33-specific deubiquitinase TRABID specifically binds K29/K33-linked diUb, and a crystal structure of this complex explains TRABID specificity and suggests a model for chain binding by TRABID. Our work uncovers linkage-specific components in the Ub system for atypical K29- and K33-linked Ub chains, providing tools to further understand these unstudied posttranslational modifications.
- Medical Research Council Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge CB2 0QH, UK.
Organizational Affiliation: