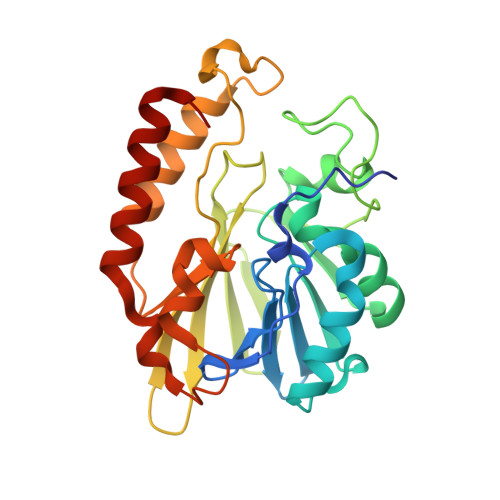
Crystal structure and kinetic analysis of the class B3 di-zinc metallo-beta-lactamase LRA-12 from an Alaskan soil metagenome.
Rodriguez, M.M., Herman, R., Ghiglione, B., Kerff, F., D'Amico Gonzalez, G., Bouillenne, F., Galleni, M., Handelsman, J., Charlier, P., Gutkind, G., Sauvage, E., Power, P.(2017) PLoS One 12: e0182043-e0182043
- PubMed: 28750094 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0182043
- Primary Citation Related Structures:
5AEB - PubMed Abstract:
We analyzed the kinetic properties of the metagenomic class B3 β-lactamase LRA-12, and determined its crystallographic structure in order to compare it with prevalent metallo-β-lactamases (MBLs) associated with clinical pathogens. We showed that LRA-12 confers extended-spectrum resistance on E. coli when expressed from recombinant clones, and the MIC values for carbapenems were similar to those observed in enterobacteria expressing plasmid-borne MBLs such as VIM, IMP or NDM. This was in agreement with the strong carbapenemase activity displayed by LRA-12, similar to GOB β-lactamases. Among the chelating agents evaluated, dipicolinic acid inhibited the enzyme more strongly than EDTA, which required pre-incubation with the enzyme to achieve measurable inhibition. Structurally, LRA-12 contains the conserved main structural features of di-zinc class B β-lactamases, and presents unique structural signatures that differentiate this enzyme from others within the family: (i) two loops (α3-β7 and β11-α5) that could influence antibiotic entrance and remodeling of the active site cavity; (ii) a voluminous catalytic cavity probably responsible for the high hydrolytic efficiency of the enzyme; (iii) the absence of disulfide bridges; (iv) a unique Gln116 at metal-binding site 1; (v) a methionine residue at position 221that replaces Cys/Ser found in other B3 β-lactamases in a predominantly hydrophobic environment, likely playing a role in protein stability. The structure of LRA-12 indicates that MBLs exist in wild microbial populations in extreme environments, or environments with low anthropic impact, and under the appropriate antibiotic selective pressure could be captured and disseminated to pathogens.
- Cátedra de Microbiología, Departamento de Microbiología, Inmunología y Biotecnología, Facultad de Farmacia y Bioquímica, Universidad de Buenos Aires, Buenos Aires, Argentina.
Organizational Affiliation: