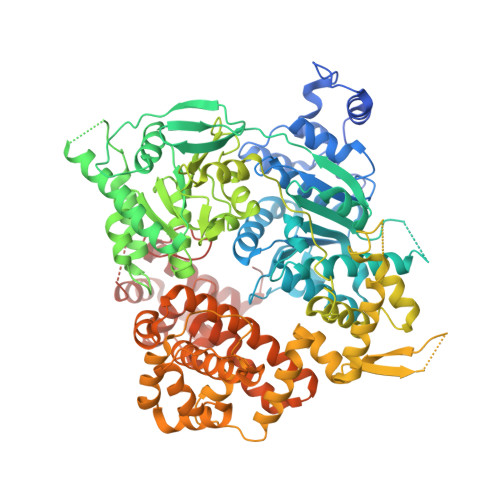
Structure of the Helicase Domain of DNA Polymerase Theta Reveals a Possible Role in the Microhomology-Mediated End-Joining Pathway.
Newman, J.A., Cooper, C.D.O., Aitkenhead, H., Gileadi, O.(2015) Structure 23: 2319
- PubMed: 26636256 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2015.10.014
- Primary Citation Related Structures:
5A9F, 5A9J, 5AGA - PubMed Abstract:
DNA polymerase theta (Polθ) has been identified as a crucial alternative non-homologous end-joining factor in mammalian cells. Polθ is upregulated in a range of cancer cell types defective in homologous recombination, and knockdown has been shown to inhibit cell survival in a subset of these, making it an attractive target for cancer treatment. We present crystal structures of the helicase domain of human Polθ in the presence and absence of bound nucleotides, and a characterization of its DNA-binding and DNA-stimulated ATPase activities. Comparisons with related helicases from the Hel308 family identify several unique features. Polθ exists as a tetramer both in the crystals and in solution. We propose a model for DNA binding to the Polθ helicase domain in the context of the Polθ tetramer, which suggests a role for the helicase domain in strand annealing of DNA templates for subsequent processing by the polymerase domain.
- Structural Genomics Consortium, University of Oxford, ORCRB, Roosevelt Drive, Oxford OX3 7DQ, UK.
Organizational Affiliation: