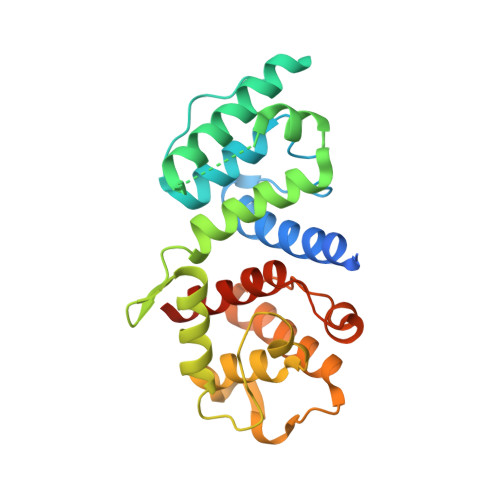
Hypertrophic Cardiomyopathy Mutations in the Calponin-Homology Domain of Actn2 Affect Actin Binding and Cardiomyocyte Z-Disc Incorporation.
Haywood, N., Wolny, M., Rogers, B., Trinh, C.H., Shuping, Y., Edwards, T.A., Peckham, M.(2016) Biochem J 473: 2485
- PubMed: 27287556 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BCJ20160421
- Primary Citation Related Structures:
5A36, 5A37, 5A38, 5A4B - PubMed Abstract:
α-Actinin-2 (ACTN2) is the only muscle isoform of α-actinin expressed in cardiac muscle. Mutations in this protein have been implicated in mild to moderate forms of hypertrophic cardiomyopathy (HCM). We have investigated the effects of two mutations identified from HCM patients, A119T and G111V, on the secondary and tertiary structure of a purified actin binding domain (ABD) of ACTN2 by circular dichroism and X-ray crystallography, and show small but distinct changes for both mutations. We also find that both mutants have reduced F-actin binding affinity, although the differences are not significant. The full length mEos2 tagged protein expressed in adult cardiomyocytes shows that both mutations additionally affect Z-disc localization and dynamic behaviour. Overall, these two mutations have small effects on structure, function and behaviour, which may contribute to a mild phenotype for this disease.
- Astbury Centre for Structural Molecular Biology and the School of Molecular and Cellular Biology, Faculty of Biological Sciences, University of Leeds, Leeds LS2 9JT, U.K.
Organizational Affiliation: