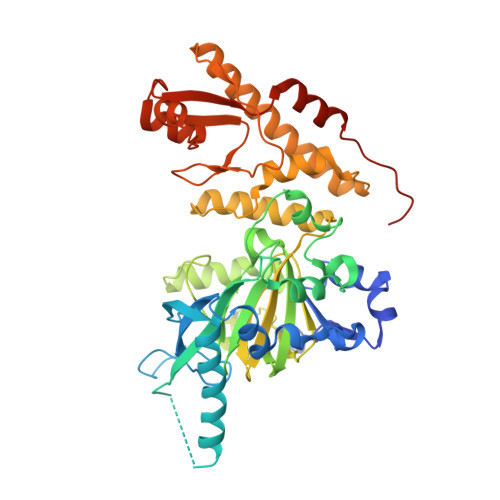
Crystal Structure of Jmjc Domain of Human Histone Demethylase Uty with S21056A
Srikannathasan, V., Gileadi, C., Johansson, C., Krojer, T., Tumber, A., von Delft, F., Arrowsmith, C.H., Bountra, C., Edwards, A., Brennan, P., Oppermann, U.To be published.