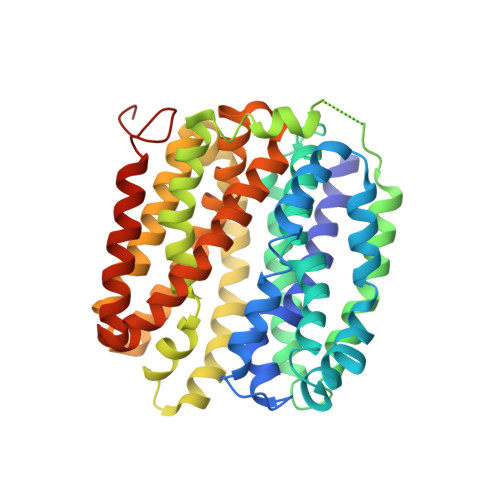
Structure of LacY with an alpha-substituted galactoside: Connecting the binding site to the protonation site.
Kumar, H., Finer-Moore, J.S., Kaback, H.R., Stroud, R.M.(2015) Proc Natl Acad Sci U S A 112: 9004-9009
- PubMed: 26157133
- DOI: https://doi.org/10.1073/pnas.1509854112
- Primary Citation Related Structures:
4ZYR - PubMed Abstract:
The X-ray crystal structure of a conformationally constrained mutant of the Escherichia coli lactose permease (the LacY double-Trp mutant Gly-46→Trp/Gly-262→Trp) with bound p-nitrophenyl-α-d-galactopyranoside (α-NPG), a high-affinity lactose analog, is described. With the exception of Glu-126 (helix IV), side chains Trp-151 (helix V), Glu-269 (helix VIII), Arg-144 (helix V), His-322 (helix X), and Asn-272 (helix VIII) interact directly with the galactopyranosyl ring of α-NPG to provide specificity, as indicated by biochemical studies and shown directly by X-ray crystallography. In contrast, Phe-20, Met-23, and Phe-27 (helix I) are within van der Waals distance of the benzyl moiety of the analog and thereby increase binding affinity nonspecifically. Thus, the specificity of LacY for sugar is determined solely by side-chain interactions with the galactopyranosyl ring, whereas affinity is increased by nonspecific hydrophobic interactions with the anomeric substituent.
- Department of Biochemistry and Biophysics, University of California, San Francisco, CA 94158;
Organizational Affiliation: