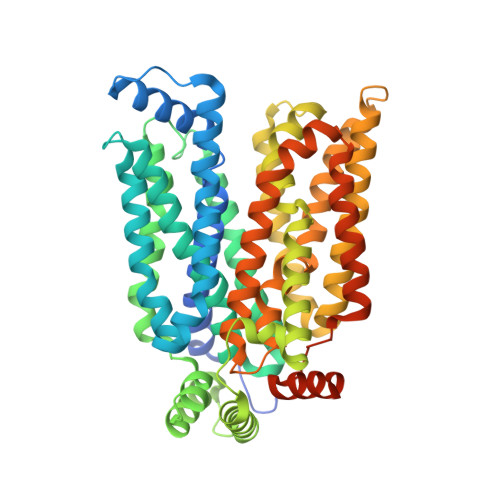
Molecular basis of ligand recognition and transport by glucose transporters
Deng, D., Sun, P.C., Yan, C.Y., Ke, M., Jiang, X., Xiong, L., Ren, W., Hirata, K., Yamamoto, M., Fan, S., Yan, N.(2015) Nature 526: 391-396
- PubMed: 26176916
- DOI: https://doi.org/10.1038/nature14655
- Primary Citation Related Structures:
4ZW9, 4ZWB, 4ZWC - PubMed Abstract:
The major facilitator superfamily glucose transporters, exemplified by human GLUT1-4, have been central to the study of solute transport. Using lipidic cubic phase crystallization and microfocus X-ray diffraction, we determined the structure of human GLUT3 in complex with D-glucose at 1.5 Å resolution in an outward-occluded conformation. The high-resolution structure allows discrimination of both α- and β-anomers of D-glucose. Two additional structures of GLUT3 bound to the exofacial inhibitor maltose were obtained at 2.6 Å in the outward-open and 2.4 Å in the outward-occluded states. In all three structures, the ligands are predominantly coordinated by polar residues from the carboxy terminal domain. Conformational transition from outward-open to outward-occluded entails a prominent local rearrangement of the extracellular part of transmembrane segment TM7. Comparison of the outward-facing GLUT3 structures with the inward-open GLUT1 provides insights into the alternating access cycle for GLUTs, whereby the C-terminal domain provides the primary substrate-binding site and the amino-terminal domain undergoes rigid-body rotation with respect to the C-terminal domain. Our studies provide an important framework for the mechanistic and kinetic understanding of GLUTs and shed light on structure-guided ligand design.
- State Key Laboratory of Membrane Biology, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: