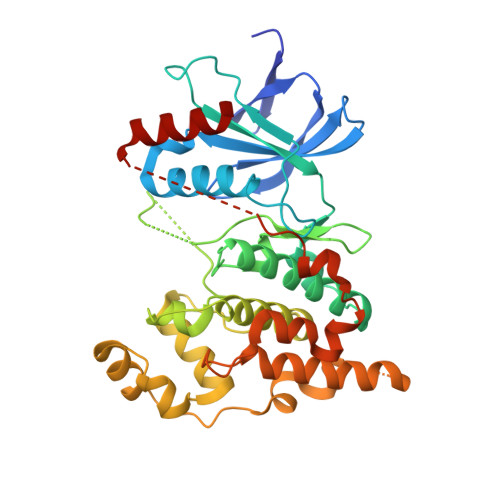
The structure of JNK3 in complex with small molecule inhibitors: structural basis for potency and selectivity.
Scapin, G., Patel, S.B., Lisnock, J., Becker, J.W., LoGrasso, P.V.(2003) Chem Biol 10: 705-712
- PubMed: 12954329
- DOI: https://doi.org/10.1016/s1074-5521(03)00159-5
- Primary Citation of Related Structures:
1PMN, 1PMU, 1PMV, 4Z9L - PubMed Abstract:
The c-Jun terminal kinases (JNKs) are members of the mitogen-activated protein (MAP) kinase family and regulate signal transduction in response to environmental stress. Activation of JNK3, a neuronal-specific isoform, has been associated with neurological damage, and as such, JNK3 may represent an attractive target for the treatment of neurological disorders. The MAP kinases share between 50% and 80% sequence identity. In order to obtain efficacious and safe compounds, it is necessary to address the issues of potency and selectivity. We report here four crystal structures of JNK3 in complex with three different classes of inhibitors. These structures provide a clear picture of the interactions that each class of compound made with the kinase. Knowledge of the atomic interactions involved in these diverse binding modes provides a platform for structure-guided modification of these compounds, or the de novo design of novel inhibitors that could satisfy the need for potency and selectivity.
- Department of Medicinal Chemistry, Merck Research Laboratories, P.O. Box 2000, Rahway, NJ 07065, USA. giovanna_scapin@merck.com
Organizational Affiliation: