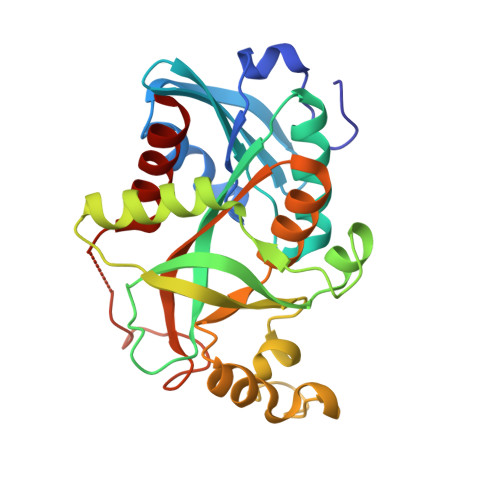
Concerted action of two subunits of the functional dimer of Shewanella oneidensis MR-1 uridine phosphorylase derived from a comparison of the C212S mutant and the wild-type enzyme.
Safonova, T.N., Mordkovich, N.N., Veiko, V.P., Okorokova, N.A., Manuvera, V.A., Dorovatovskii, P.V., Popov, V.O., Polyakov, K.M.(2016) Acta Crystallogr D Struct Biol 72: 203-210
- PubMed: 26894668
- DOI: https://doi.org/10.1107/S2059798315024353
- Primary Citation Related Structures:
4YJK - PubMed Abstract:
Uridine phosphorylase (UP; EC 2.4.2.3), a key enzyme in the pyrimidine-salvage pathway, catalyzes the reversible phosphorolysis of uridine to uracil and ribose 1-phosphate. The structure of the C212S mutant of uridine phosphorylase from the facultatively aerobic Gram-negative γ-proteobacterium Shewanella oneidensis MR-1 (SoUP) was determined at 1.68 Å resolution. A comparison of the structures of the mutant and the wild-type enzyme showed that one dimer in the mutant hexamer differs from all other dimers in the mutant and wild-type SoUP (both in the free form and in complex with uridine). The key difference is the `maximum open' state of one of the subunits comprising this dimer, which has not been observed previously for uridine phosphorylases. Some conformational features of the SoUP dimer that provide access of the substrate into the active site are revealed. The binding of the substrate was shown to require the concerted action of two subunits of the dimer. The changes in the three-dimensional structure induced by the C212S mutation account for the lower affinity of the mutant for inorganic phosphate, while the affinity for uridine remains unchanged.
- Bach Institute of Biochemistry, Russian Academy of Sciences, 33 Leninskii Ave., Moscow 119071, Russian Federation.
Organizational Affiliation: