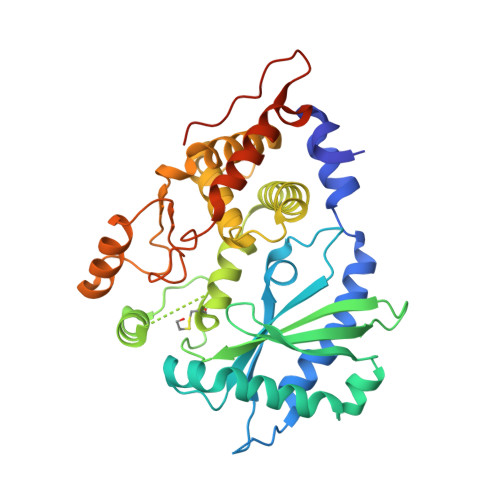
Structural and functional analysis reveals that human OASL binds dsRNA to enhance RIG-I signaling.
Ibsen, M.S., Gad, H.H., Andersen, L.L., Hornung, V., Julkunen, I., Sarkar, S.N., Hartmann, R.(2015) Nucleic Acids Res 43: 5236-5248
- PubMed: 25925578
- DOI: https://doi.org/10.1093/nar/gkv389
- Primary Citation of Related Structures:
4XQ7 - PubMed Abstract:
The oligoadenylate synthetase (OAS) enzymes are cytoplasmic dsRNA sensors belonging to the antiviral innate immune system. Upon binding to viral dsRNA, the OAS enzymes synthesize 2'-5' linked oligoadenylates (2-5As) that initiate an RNA decay pathway to impair viral replication. The human OAS-like (OASL) protein, however, does not harbor the catalytic activity required for synthesizing 2-5As and differs from the other human OAS family members by having two C-terminal ubiquitin-like domains. In spite of its lack of enzymatic activity, human OASL possesses antiviral activity. It was recently demonstrated that the ubiquitin-like domains of OASL could substitute for K63-linked poly-ubiquitin and interact with the CARDs of RIG-I and thereby enhance RIG-I signaling. However, the role of the OAS-like domain of OASL remains unclear. Here we present the crystal structure of the OAS-like domain, which shows a striking similarity with activated OAS1. Furthermore, the structure of the OAS-like domain shows that OASL has a dsRNA binding groove. We demonstrate that the OAS-like domain can bind dsRNA and that mutating key residues in the dsRNA binding site is detrimental to the RIG-I signaling enhancement. Hence, binding to dsRNA is an important feature of OASL that is required for enhancing RIG-I signaling.
- Centre for Structural Biology, Department of Molecular Biology and Genetics, Aarhus University, 8000 Aarhus C, Denmark.
Organizational Affiliation: