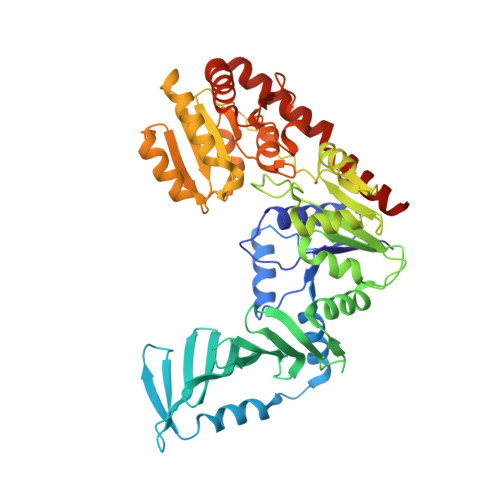
Structure and mechanism of Staphylococcus aureus TarM, the wall teichoic acid alpha-glycosyltransferase.
Sobhanifar, S., Worrall, L.J., Gruninger, R.J., Wasney, G.A., Blaukopf, M., Baumann, L., Lameignere, E., Solomonson, M., Brown, E.D., Withers, S.G., Strynadka, N.C.(2015) Proc Natl Acad Sci U S A 112: E576-E585
- PubMed: 25624472
- DOI: https://doi.org/10.1073/pnas.1418084112
- Primary Citation Related Structures:
4X6L, 4X7M, 4X7P, 4X7R - PubMed Abstract:
Unique to Gram-positive bacteria, wall teichoic acids are anionic glycopolymers cross-stitched to a thick layer of peptidoglycan. The polyol phosphate subunits of these glycopolymers are decorated with GlcNAc sugars that are involved in phage binding, genetic exchange, host antibody response, resistance, and virulence. The search for the enzymes responsible for GlcNAcylation in Staphylococcus aureus has recently identified TarM and TarS with respective α- and β-(1-4) glycosyltransferase activities. The stereochemistry of the GlcNAc attachment is important in balancing biological processes, such that the interplay of TarM and TarS is likely important for bacterial pathogenicity and survival. Here we present the crystal structure of TarM in an unusual ternary-like complex consisting of a polymeric acceptor substrate analog, UDP from a hydrolyzed donor, and an α-glyceryl-GlcNAc product formed in situ. These structures support an internal nucleophilic substitution-like mechanism, lend new mechanistic insight into the glycosylation of glycopolymers, and reveal a trimerization domain with a likely role in acceptor substrate scaffolding.
- Department of Biochemistry and Center for Blood Research, University of British Columbia, Vancouver, BC, Canada V6T 1Z3;
Organizational Affiliation: