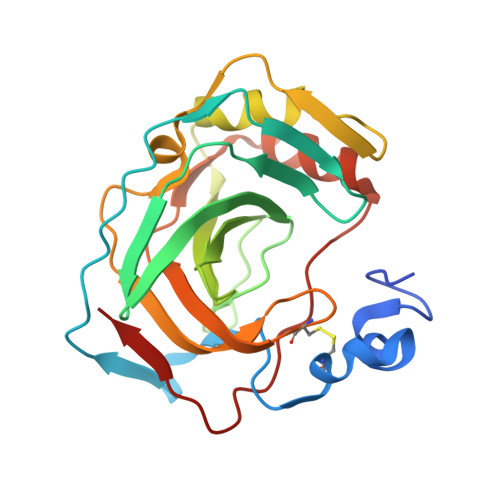
Crystal structure of the most catalytically effective carbonic anhydrase enzyme known, SazCA from the thermophilic bacterium Sulfurihydrogenibium azorense.
De Simone, G., Monti, S.M., Alterio, V., Buonanno, M., De Luca, V., Rossi, M., Carginale, V., Supuran, C.T., Capasso, C., Di Fiore, A.(2015) Bioorg Med Chem Lett 25: 2002-2006
- PubMed: 25817590
- DOI: https://doi.org/10.1016/j.bmcl.2015.02.068
- Primary Citation Related Structures:
4X5S - PubMed Abstract:
Two thermostable α-carbonic anhydrases (α-CAs) isolated from thermophilic Sulfurihydrogenibium spp., namely SspCA (from S. yellowstonensis) and SazCA (from S. azorense), were shown in a previous work to possess interesting complementary properties. SspCA was shown to have an exceptional thermal stability, whereas SazCA demonstrated to be the most active α-CA known to date for the CO2 hydration reaction. Here we report the crystallographic structure of SazCA and the identification of the structural features responsible for its high catalytic activity, by comparing it with SspCA structure. These data are of relevance for the design of engineered proteins showing higher stability and catalytic activity than other α-CAs known to date.
- Istituto di Biostrutture e Bioimmagini-CNR, via Mezzocannone 16, 80134 Napoli, Italy. Electronic address: gdesimon@unina.it.
Organizational Affiliation: