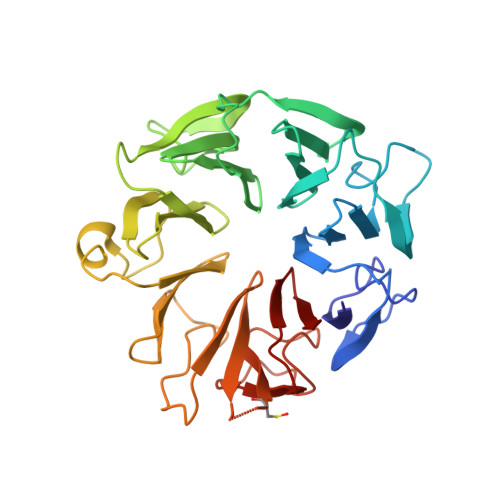
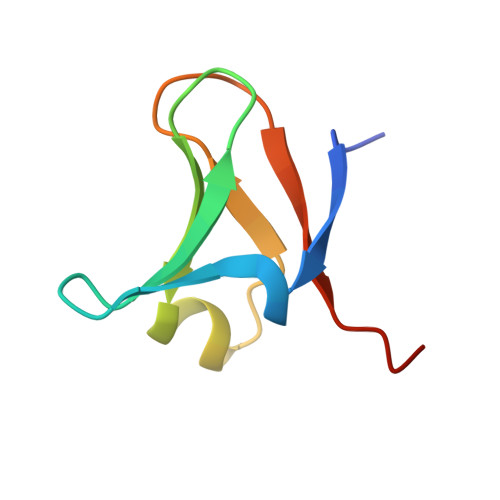
Structure of the Elongator cofactor complex Kti11/Kti13 provides insight into the role of Kti13 in Elongator-dependent tRNA modification.
Kolaj-Robin, O., McEwen, A.G., Cavarelli, J., Seraphin, B.(2015) FEBS J 282: 819-833
- PubMed: 25604895 Search on PubMed
- DOI: https://doi.org/10.1111/febs.13199
- Primary Citation Related Structures:
4X33 - PubMed Abstract:
Modification of wobble uridines of many eukaryotic tRNAs requires the Elongator complex, a highly conserved six-subunit eukaryotic protein assembly, as well as the Killer toxin-insensitive (Kti) proteins 11-14. Kti11 was additionally shown to be implicated in the biosynthesis of diphthamide, a post-translationally modified histidine of translation elongation factor 2. Recent data indicate that iron-bearing Kti11 functions as an electron donor to the [4Fe-4S] cluster of radical S-Adenosylmethionine enzymes, triggering the subsequent radical reaction. We show here that recombinant yeast Kti11 forms a stable 1 : 1 complex with Kti13. To obtain insights into the function of this heterodimer, the Kti11/Kti13 complex was purified to homogeneity, crystallized, and its structure determined at 1.45 Å resolution. The importance of several residues mediating complex formation was confirmed by mutagenesis. Kti13 adopts a fold characteristic of RCC1-like proteins. The seven-bladed β-propeller consists of a unique mixture of four- and three-stranded blades. In the complex, Kti13 orients Kti11 and restricts access to its electron-carrying iron atom, constraining the electron transfer capacity of Kti11. Based on these findings, we propose a role for Kti13, and discuss the possible functional implications of complex formation. Structural data have been submitted to the Protein Data Bank under accession number 4X33.
- Equipe Labellisée La Ligue, Institut de Génétique et de Biologie Moléculaire et Cellulaire, Centre National de Recherche Scientifique UMR 7104/Institut National de Santé et de Recherche Médicale U964/Université de Strasbourg, Illkirch, France.
Organizational Affiliation: