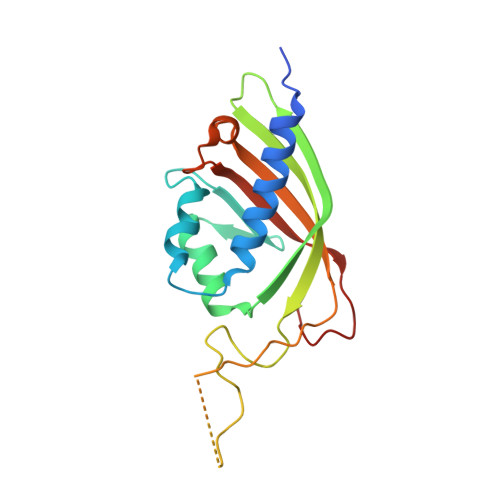
Domain organization within the nuclear export factor Mex67:Mtr2 generates an extended mRNA binding surface.
Aibara, S., Valkov, E., Lamers, M., Stewart, M.(2015) Nucleic Acids Res 43: 1927-1936
- PubMed: 25618852 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkv030
- Primary Citation Related Structures:
4WWU - PubMed Abstract:
The Mex67:Mtr2 complex is the principal yeast nuclear export factor for bulk mRNA and also contributes to ribosomal subunit export. Mex67 is a modular protein constructed from four domains (RRM, LRR, NTF2-like and UBA) that have been thought to be joined by flexible linkers like beads on a string, with the RRM and LRR domains binding RNAs and the NTF2-like and UBA domains binding FG-nucleoporins to facilitate movement through nuclear pores. Here, we show that the NTF2-like domain from Saccharomyces cerevisiae Mex67:Mtr2 also contributes to RNA binding. Moreover, the 3.3 Å resolution crystal structure of the Mex67(ΔUBA):Mtr2 complex, supplemented with small angle X-ray scattering data, indicated that the LRR domain has a defined spatial relationship to the Mex67(NTF2L):Mtr2 region. Conversely, the RRM domain and especially the UBA domain are more mobile. The conformation assumed by the LRR and NTF2-like domains results in clusters of positively-charged residues on each becoming arranged to form a continuous interface for binding RNA on the opposite side of the complex to the region that interacts with FG-nucleoporins to facilitate passage through nuclear pores.
- MRC Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge Biomedical Campus, Cambridge CB2 0QH, UK.
Organizational Affiliation: