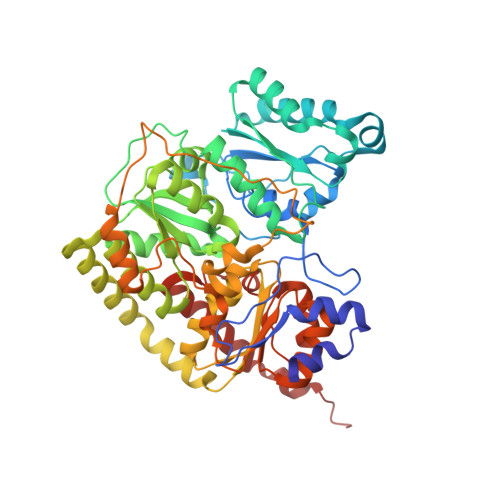
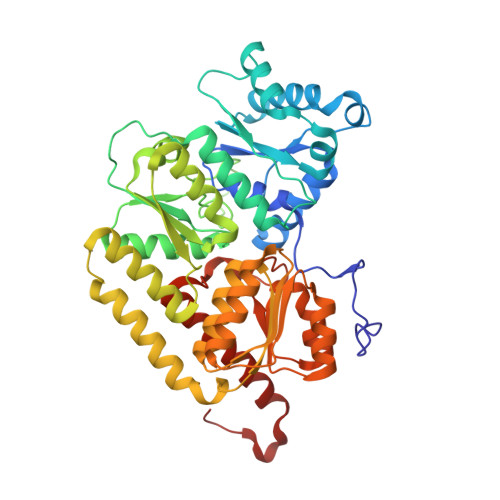
Substrate Pathways in the Nitrogenase MoFe Protein by Experimental Identification of Small Molecule Binding Sites.
Morrison, C.N., Hoy, J.A., Zhang, L., Einsle, O., Rees, D.C.(2015) Biochemistry 54: 2052-2060
- PubMed: 25710326 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi501313k
- Primary Citation Related Structures:
4WN9, 4WNA - PubMed Abstract:
In the nitrogenase molybdenum-iron (MoFe) protein, we have identified five potential substrate access pathways from the protein surface to the FeMo-cofactor (the active site) or the P-cluster using experimental structures of Xe pressurized into MoFe protein crystals from Azotobacter vinelandii and Clostridium pasteurianum. Additionally, all published structures of the MoFe protein, including those from Klebsiella pneumoniae, were analyzed for the presence of nonwater, small molecules bound to the protein interior. Each pathway is based on identification of plausible routes from buried small molecule binding sites to both the protein surface and a metallocluster. Of these five pathways, two have been previously suggested as substrate access pathways. While the small molecule binding sites are not conserved among the three species of MoFe protein, residues lining the pathways are generally conserved, indicating that the proposed pathways may be accessible in all three species. These observations imply that there is unlikely a unique pathway utilized for substrate access from the protein surface to the active site; however, there may be preferred pathways such as those described here.
- †Division of Chemistry and Chemical Engineering, California Institute of Technology 114-96, Pasadena, California 91125, United States.
Organizational Affiliation: