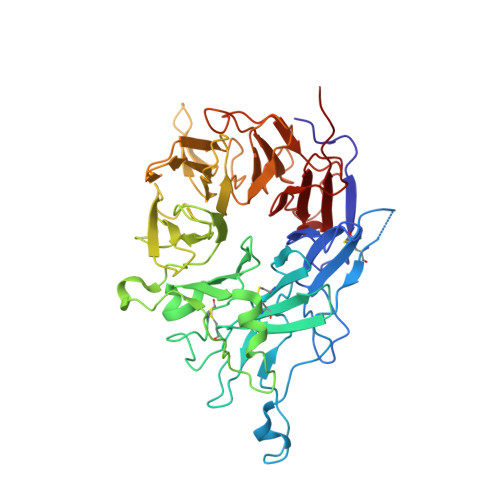
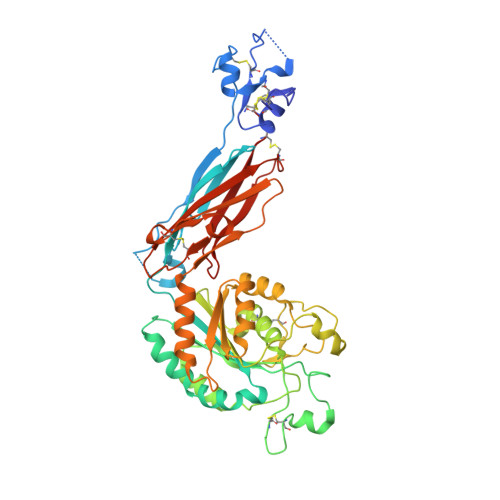
Metal ion and ligand binding of integrin alpha 5 beta 1.
Xia, W., Springer, T.A.(2014) Proc Natl Acad Sci U S A 111: 17863-17868
- PubMed: 25475857 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1420645111
- Primary Citation Related Structures:
4WJK, 4WK0, 4WK2, 4WK4 - PubMed Abstract:
Integrin α5β1 binds to an Arg-Gly-Asp (RGD) motif in its ligand fibronectin. We report high-resolution crystal structures of a four-domain α5β1 headpiece fragment, alone or with RGD peptides soaked into crystals, and RGD peptide affinity measurements. The headpiece crystallizes in a closed conformation essentially identical to that seen previously for α5β1 complexed with a Fab that allosterically inhibits ligand binding by stabilizing the closed conformation. Soaking experiments show that binding of cyclic RGD peptide with 20-fold higher affinity than a linear RGD peptide induces conformational change in the β1-subunit βI domain to a state that is intermediate between closed (low affinity) and open (high affinity). In contrast, binding of a linear RGD peptide induces no shape shifting. However, linear peptide binding induces shape shifting when Ca(2+) is depleted during soaking. Ca(2+) bound to the adjacent to metal ion-dependent adhesion site (ADMIDAS), at the locus of shape shifting, moves and decreases in occupancy, correlating with an increase in affinity for RGD measured when Ca(2+) is depleted. The results directly demonstrate that Ca(2+) binding to the ADMIDAS stabilizes integrins in the low-affinity, closed conformation. Comparisons in affinity between four-domain and six-domain headpiece constructs suggest that flexible integrin leg domains contribute to conformational equilibria. High-resolution views of the hybrid domain interface with the plexin-semaphorin-integrin (PSI) domain in different orientations show a ball-and-socket joint with a hybrid domain Arg side chain that rocks in a PSI domain socket lined with carbonyl oxygens.
- Program in Cellular and Molecular Medicine, Boston Children's Hospital, Boston, MA 02115; and Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115.
Organizational Affiliation: