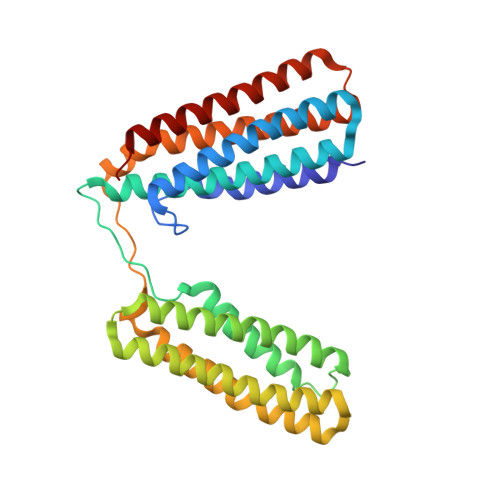
Structural and Mechanistic Insights into the Recruitment of Talin by RIAM in Integrin Signaling.
Chang, Y.C., Zhang, H., Franco-Barraza, J., Brennan, M.L., Patel, T., Cukierman, E., Wu, J.(2014) Structure 22: 1810-1820
- PubMed: 25465129 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2014.09.020
- Primary Citation Related Structures:
4W8P - PubMed Abstract:
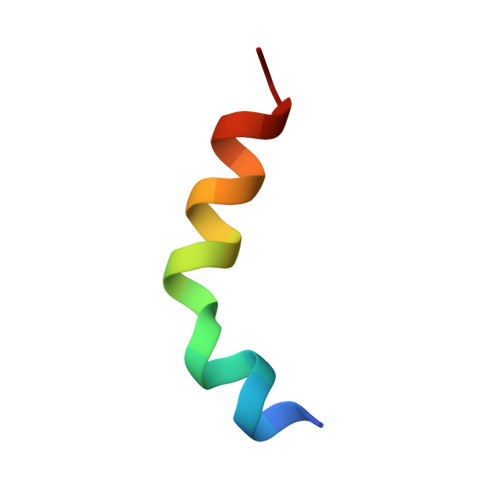
Plasma membrane (PM)-bound GTPase Rap1 recruits the Rap1-interacting-adaptor-molecule (RIAM), which in turn recruits talin to bind and activate integrins. However, it is unclear how RIAM recruits talin and why its close homolog lamellipodin does not. Here, we report that, although RIAM possesses two talin-binding sites (TBS1 and TBS2), only TBS1 is capable of recruiting cytoplasmic talin to the PM, and the R8 domain is the strongest binding site in talin. Crystal structure of an R7R8:TBS1 complex reveals an unexpected kink in the TBS1 helix that is not shared in the homologous region of lamellipodin. This kinked helix conformation is required for the colocalization of RIAM and talin at the PM and proper activation of integrin. Our findings provide the structural and mechanistic insight into talin recruitment by RIAM that underlies integrin activation and explain the differential functions of the otherwise highly homologous RIAM and lamellipodin in integrin signaling.
- Developmental Therapeutics Program, Fox Chase Cancer Center, 333 Cottman Avenue, Philadelphia, PA 19111, USA.
Organizational Affiliation: