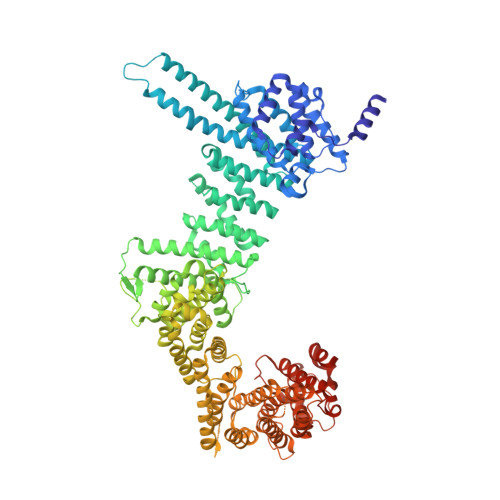
Structure and Function of Cohesins Scc3/Sa Regulatory Subunit
Roig, M.B., Lowe, J., Chan, K.L., Beckouet, F., Metson, J., Nasmyth, K.(2014) FEBS Lett 588: 3692
- PubMed: 25171859
- DOI: https://doi.org/10.1016/j.febslet.2014.08.015
- Primary Citation Related Structures:
4UVJ, 4UVK - PubMed Abstract:
Sister chromatid cohesion involves entrapment of sister DNAs by a cohesin ring created through association of a kleisin subunit (Scc1) with ATPase heads of Smc1/Smc3 heterodimers. Cohesin's association with chromatin involves subunits recruited by Scc1: Wapl, Pds5, and Scc3/SA, in addition to Scc2/4 loading complex. Unlike Pds5, Wapl, and Scc2/4, Scc3s are encoded by all eukaryotic genomes. Here, a crystal structure of Scc3 reveals a hook-shaped protein composed of tandem α helices. Its N-terminal domain contains a conserved and essential surface (CES) present even in organisms lacking Pds5, Wapl, and Scc2/4, while its C-terminal domain binds a section of the kleisin Scc1. Scc3 turns over in G2/M while maintaining cohesin's association with chromosomes and it promotes de-acetylation of Smc3 upon Scc1 cleavage.
- Department of Biochemistry, University of Oxford, Oxford OX1 3QU, United Kingdom.
Organizational Affiliation: