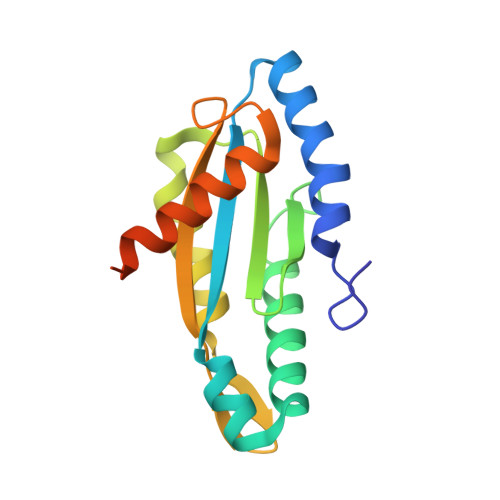
Structure of a Diguanylate Cyclase from Thermotoga Maritima: Insights Into Activation, Feedback Inhibition and Thermostability
Deepthi, A., Liew, C.W., Liang, Z.X., Kunchithapadam, S., Lescar, J.(2014) PLoS One 9: 10912
- PubMed: 25360685 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0110912
- Primary Citation Related Structures:
4URG, 4URQ, 4URS - PubMed Abstract:
Large-scale production of bis-3'-5'-cyclic-di-GMP (c-di-GMP) would facilitate biological studies of numerous bacterial signaling pathways and phenotypes controlled by this second messenger molecule, such as virulence and biofilm formation. C-di-GMP constitutes also a potentially interesting molecule as a vaccine adjuvant. Even though chemical synthesis of c-di-GMP can be done, the yields are incompatible with mass-production. tDGC, a stand-alone diguanylate cyclase (DGC or GGDEF domain) from Thermotoga maritima, enables the robust enzymatic production of large quantities of c-di-GMP. To understand the structural correlates of tDGC thermostability, its catalytic mechanism and feedback inhibition, we determined structures of an active-like dimeric conformation with both active (A) sites facing each other and of an inactive dimeric conformation, locked by c-di-GMP bound at the inhibitory (I) site. We also report the structure of a single mutant of tDGC, with the R158A mutation at the I-site, abolishing product inhibition and unproductive dimerization. A comparison with structurally characterized DGC homologues from mesophiles reveals the presence of a higher number of salt bridges in the hyperthermophile enzyme tDGC. Denaturation experiments of mutants disrupting in turn each of the salt bridges unique to tDGC identified three salt-bridges critical to confer thermostability.
- Department of Biological Sciences, National University of Singapore, Singapore, Singapore.
Organizational Affiliation: