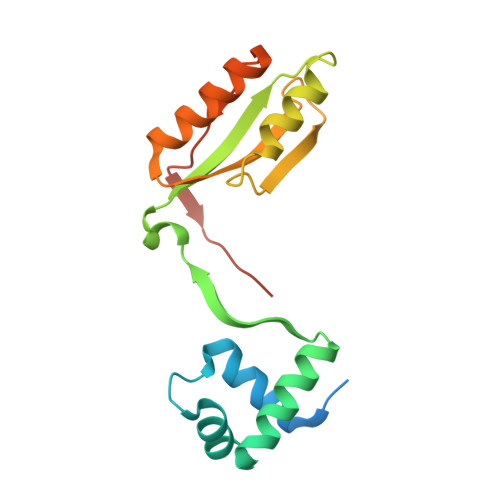
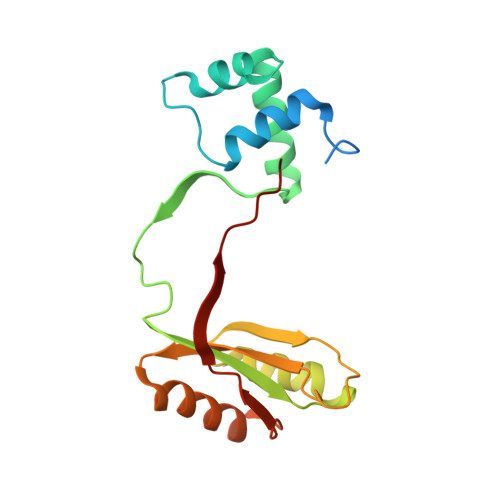
The Structure, Function and Properties of Sirohaem Decarboxylase - an Enzyme with Structural Homology to a Transcription Factor Family that is Part of the Alternative Haem Biosynthesis Pathway.
Palmer, D.J., Schroeder, S., Lawrence, A.D., Deery, E., Lobo, S.A., Saraiva, L.M., Mclean, K.J., Munro, A.W., Ferguson, S.J., Pickersgill, R.W., Brown, D.G., Warren, M.J.(2014) Mol Microbiol 93: 247
- PubMed: 24865947
- DOI: https://doi.org/10.1111/mmi.12656
- Primary Citation Related Structures:
4CZD, 4UN1 - PubMed Abstract:
Some bacteria and archaea synthesize haem by an alternative pathway, which involves the sequestration of sirohaem as a metabolic intermediate rather than as a prosthetic group. Along this pathway the two acetic acid side-chains attached to C12 and C18 are decarboxylated by sirohaem decarboxylase, a heterodimeric enzyme composed of AhbA and AhbB, to give didecarboxysirohaem. Further modifications catalysed by two related radical SAM enzymes, AhbC and AhbD, transform didecarboxysirohaem into Fe-coproporphyrin III and haem respectively. The characterization of sirohaem decarboxylase is reported in molecular detail. Recombinant versions of Desulfovibrio desulfuricans, Desulfovibrio vulgaris and Methanosarcina barkeri AhbA/B have been produced and their physical properties compared. The D. vulgaris and M. barkeri enzyme complexes both copurify with haem, whose redox state influences the activity of the latter. The kinetic parameters of the D. desulfuricans enzyme have been determined, the enzyme crystallized and its structure has been elucidated. The topology of the enzyme reveals that it shares a structural similarity to the AsnC/Lrp family of transcription factors. The active site is formed in the cavity between the two subunits and a AhbA/B-product complex with didecarboxysirohaem has been obtained. A mechanism for the decarboxylation of the kinetically stable carboxyl groups is proposed.
- School of Biosciences, University of Kent, Giles Lane, Canterbury, Kent, CT2 7NJ, UK.
Organizational Affiliation: