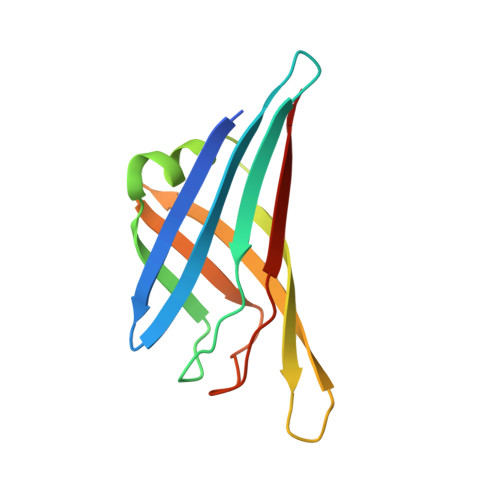
Crystal structure of the fibre head domain of bovine adenovirus 4, a ruminant atadenovirus.
Nguyen, T.H., Vidovszky, M.Z., Ballmann, M.Z., Sanz-Gaitero, M., Singh, A.K., Harrach, B., Benko, M., van Raaij, M.J.(2015) Virol J 12: 81-81
- PubMed: 25994880 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/s12985-015-0309-1
- Primary Citation Related Structures:
4UE0 - PubMed Abstract:
In adenoviruses, primary host cell recognition is generally performed by the head domains of their homo-trimeric fibre proteins. This first interaction is reversible. A secondary, irreversible interaction subsequently takes place via other adenovirus capsid proteins and leads to a productive infection. Although many fibre head structures are known for human mastadenoviruses, not many animal adenovirus fibre head structures have been determined, especially not from those belonging to adenovirus genera other than Mastadenovirus. We constructed an expression vector for the fibre head domain from a ruminant atadenovirus, bovine adenovirus 4 (BAdV-4), consisting of amino acids 414-535, expressed the protein in Escherichia coli, purified it by metal affinity and cation exchange chromatography and crystallized it. The structure was solved using single isomorphous replacement plus anomalous dispersion of a mercury derivative and refined against native data that extended to 1.2 Å resolution. Like in other adenoviruses, the BAdV-4 fibre head monomer contains a beta-sandwich consisting of ABCJ and GHID sheets. The topology is identical to the fibre head of the other studied atadenovirus, snake adenovirus 1 (SnAdV-1), including the alpha-helix in the DG-loop, despite of them having a sequence identity of only 15 %. There are also differences which may have implications for ligand binding. Beta-strands G and H are longer and differences in several surface-loops and surface charge are observed. Chimeric adenovirus fibres have been used to retarget adenovirus-based anti-cancer and gene therapy vectors. Ovine adenovirus 7 (OAdV-7), another ruminant atadenovirus, is intensively tested as a basis for such a vector. Here, we present the high-resolution atomic structure of the BAdV-4 fibre head domain, the second atadenovirus fibre head structure known and the first of an atadenovirus that infects a mammalian host. Future research should focus on the receptor-binding properties of these fibre head domains.
- Departamento de Estructura de Macromoleculas, Centro Nacional de Biotecnologia (CNB-CSIC), calle Darwin 3, 28049, Madrid, Spain. th.nguyen@cnb.csic.es.
Organizational Affiliation: