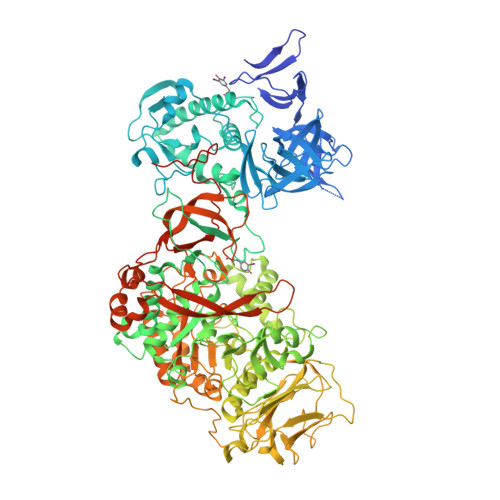
Structural Insights into the Carbohydrate Binding Ability of an alpha-(12) Branching Sucrase from Glycoside Hydrolase Family 70.
Brison, Y., Malbert, Y., Czaplicki, G., Mourey, L., Remaud-Simeon, M., Tranier, S.(2016) J Biological Chem 291: 7527-7540
- PubMed: 26865636
- DOI: https://doi.org/10.1074/jbc.M115.688796
- Primary Citation Related Structures:
4TTU, 4TVC, 4TVD - PubMed Abstract:
The α-(1→2) branching sucrase ΔN123-GBD-CD2 is a transglucosylase belonging to glycoside hydrolase family 70 (GH70) that catalyzes the transfer ofd-glucosyl units from sucroseto dextrans or gluco-oligosaccharides via the formation of α-(1→2) glucosidic linkages. The first structures of ΔN123-GBD-CD2 in complex withd-glucose, isomaltosyl, or isomaltotriosyl residues were solved. The glucose complex revealed three glucose-binding sites in the catalytic gorge and six additional binding sites at the surface of domains B, IV, and V. Soaking with isomaltotriose or gluco-oligosaccharides led to structures in which isomaltosyl or isomaltotriosyl residues were found in glucan binding pockets located in domain V. One aromatic residue is systematically identified at the bottom of these pockets in stacking interaction with one glucosyl moiety. The carbohydrate is also maintained by a network of hydrogen bonds and van der Waals interactions. The sequence of these binding pockets is conserved and repeatedly present in domain V of several GH70 glucansucrases known to bind α-glucans. These findings provide the first structural evidence of the molecular interaction occurring between isomalto-oligosaccharides and domain V of the GH70 enzymes.
- the INRA, UMR792 Ingénierie des Systèmes Biologiques et des Procédés, F-31400 Toulouse.
Organizational Affiliation: