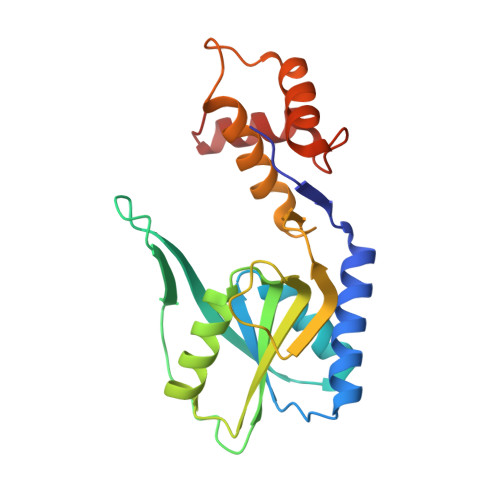
Adenine Phosphoribosyltransferase from Sulfolobus solfataricus Is an Enzyme with Unusual Kinetic Properties and a Crystal Structure that Suggests It Evolved from a 6-Oxopurine Phosphoribosyltransferase.
Jensen, K.F., Hansen, M.R., Jensen, K.S., Christoffersen, S., Poulsen, J.C., Mlgaard, A., Kadziola, A.(2015) Biochemistry 54: 2323-2334
- PubMed: 25790177 Search on PubMed
- DOI: https://doi.org/10.1021/bi501334m
- Primary Citation Related Structures:
4TRB, 4TRC, 4TS5, 4TS7 - PubMed Abstract:
The adenine phosphoribosyltransferase (APRTase) encoded by the open reading frame SSO2342 of Sulfolobus solfataricus P2 was subjected to crystallographic, kinetic, and ligand binding analyses. The enzyme forms dimers in solution and in the crystals, and binds one molecule of the reactants 5-phosphoribosyl-α-1-pyrophosphate (PRPP) and adenine or the product adenosine monophosphate (AMP) or the inhibitor adenosine diphosphate (ADP) in each active site. The individual subunit adopts an overall structure that resembles a 6-oxopurine phosphoribosyltransferase (PRTase) more than known APRTases implying that APRT functionality in Crenarchaeotae has its evolutionary origin in this family of PRTases. Only the N-terminal two-thirds of the polypeptide chain folds as a traditional type I PRTase with a five-stranded β-sheet surrounded by helices. The C-terminal third adopts an unusual three-helix bundle structure that together with the nucleobase-binding loop undergoes a conformational change upon binding of adenine and phosphate resulting in a slight contraction of the active site. The inhibitor ADP binds like the product AMP with both the α- and β-phosphates occupying the 5'-phosphoribosyl binding site. The enzyme shows activity over a wide pH range, and the kinetic and ligand binding properties depend on both pH and the presence/absence of phosphate in the buffers. A slow hydrolysis of PRPP to ribose 5-phosphate and pyrophosphate, catalyzed by the enzyme, may be facilitated by elements in the C-terminal three-helix bundle part of the protein.
- †University of Copenhagen, Department of Biology, Ole Maaløes Vej 5, DK-2200 Copenhagen N, Denmark.
Organizational Affiliation: