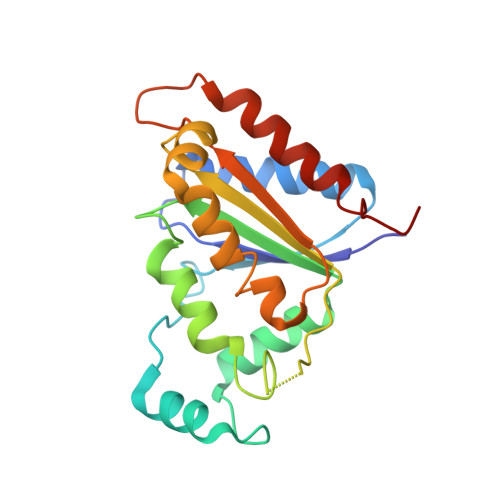
A novel cytosolic NADH:quinone oxidoreductase from Methanothermobacter marburgensis.
Ullmann, E., Tan, T.C., Gundinger, T., Herwig, C., Divne, C., Spadiut, O.(2014) Biosci Rep 34: e00167-e00167
- PubMed: 25372605
- DOI: https://doi.org/10.1042/BSR20140143
- Primary Citation Related Structures:
4R81 - PubMed Abstract:
Methanothermobacter marburgensis is a strictly anaerobic, thermophilic methanogenic archaeon that uses methanogenesis to convert H2 and CO2 to energy. M. marburgensis is one of the best-studied methanogens, and all genes required for methanogenic metabolism have been identified. Nonetheless, the present study describes a gene (Gene ID 9704440) coding for a putative quinone oxidoreductase that has not yet been identified as part of the metabolic machinery. The gene product, MmNQO, was successfully expressed, purified and characterized biochemically, as well as structurally. MmNQO was identified as a flavin-dependent NADH:quinone oxidoreductase with the capacity to oxidize NADH in the presence of a wide range of electron acceptors, whereas NADPH was oxidized with only three acceptors. The 1.50 Å crystal structure of MmNQO features a homodimeric enzyme where each monomer comprises 196 residues folding into flavodoxin-like α/β domains with non-covalently bound FMN (flavin mononucleotide). The closest structural homologue is the modulator of drug activity B from Streptococcus mutans with 1.6 Å root-mean-square deviation on 161 Cα atoms and 28% amino-acid sequence identity. The low similarity at sequence and structural level suggests that MmNQO is unique among NADH:quinone oxidoreductases characterized to date. Based on preliminary bioreactor experiments, MmNQO could provide a useful tool to prevent overflow metabolism in applications that require cells with high energy demand.
- *Vienna University of Technology, Institute of Chemical Engineering, Research Area Biochemical Engineering, Vienna, Austria.
Organizational Affiliation: