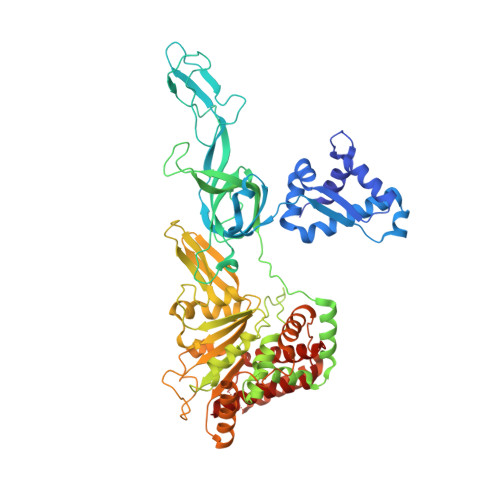
Analysis of the crystal structure of an active MCM hexamer.
Miller, J.M., Arachea, B.T., Epling, L.B., Enemark, E.J.(2014) Elife 3: e03433-e03433
- PubMed: 25262915
- DOI: https://doi.org/10.7554/eLife.03433
- Primary Citation Related Structures:
4R7Y, 4R7Z - PubMed Abstract:
In a previous Research article (Froelich et al., 2014), we suggested an MCM helicase activation mechanism, but were limited in discussing the ATPase domain because it was absent from the crystal structure. Here we present the crystal structure of a nearly full-length MCM hexamer that is helicase-active and thus has all features essential for unwinding DNA. The structure is a chimera of Sulfolobus solfataricus N-terminal domain and Pyrococcus furiosus ATPase domain. We discuss three major findings: 1) a novel conformation for the A-subdomain that could play a role in MCM regulation; 2) interaction of a universally conserved glutamine in the N-terminal Allosteric Communication Loop with the AAA+ domain helix-2-insert (h2i); and 3) a recessed binding pocket for the MCM ssDNA-binding motif influenced by the h2i. We suggest that during helicase activation, the h2i clamps down on the leading strand to facilitate strand retention and regulate ATP hydrolysis.
- Department of Structural Biology, St Jude Children's Research Hospital, Memphis, United States.
Organizational Affiliation: