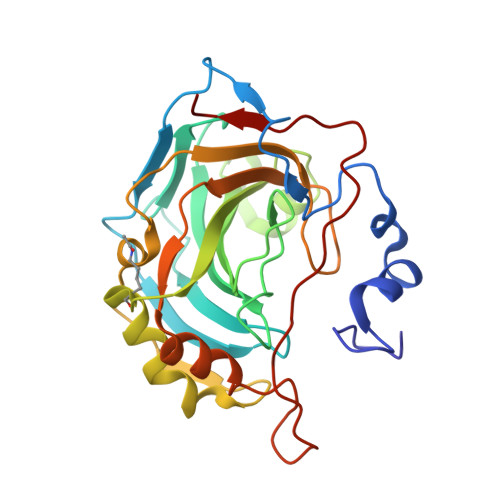
Structural Insights into Carbonic Anhydrase IX Isoform Specificity of Carbohydrate-Based Sulfamates.
Moeker, J., Mahon, B.P., Bornaghi, L.F., Vullo, D., Supuran, C.T., McKenna, R., Poulsen, S.A.(2014) J Med Chem 57: 8635-8645
- PubMed: 25254302 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jm5012935
- Primary Citation Related Structures:
4R59, 4R5A, 4R5B - PubMed Abstract:
Carbonic anhydrase IX (CA IX) is an extracellular transmembrane homodimeric zinc metalloenzyme that has been validated as a prognostic marker and therapeutic target for several types of aggressive cancers. CA IX shares a close homology with other CA isoforms, making the design of CA IX isoform selective inhibitors challenging. In this paper, we describe the development of a new class of CA IX inhibitors that comprise a sulfamate as the zinc binding group, a variable linker, and a carbohydrate "tail" moiety. Seven compounds inhibited CA IX with low nM Ki values of 1-2 nM and also exhibited permeability profiles to preferentially target the binding of extracellular CA IX over cytosolic CAs. The crystal structures of two of these compounds in complex with a CA IX-mimic (a variant of CA II, with active site residues that mimic CA IX) and one compound in complex with CA II have been determined to 1.7 Å resolution or better and demonstrate a selective mechanism of binding between the hydrophilic and hydrophobic pockets of CA IX versus CA II. These compounds present promising candidates for anti-CA IX drugs and the treatment for several aggressive cancer types.
- Eskitis Institute for Drug Discovery, Griffith University , Nathan, Queensland 4111, Australia.
Organizational Affiliation: