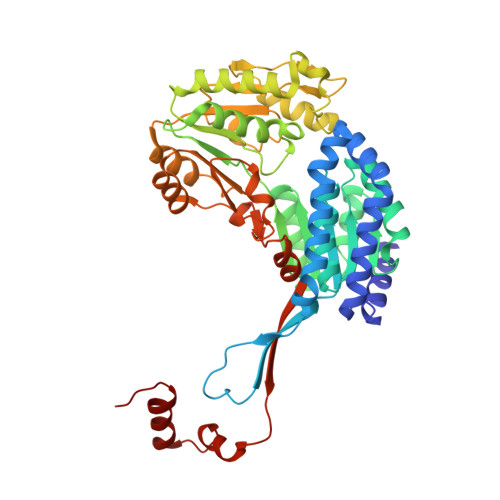
A gatekeeper helix determines the substrate specificity of Sjogren-Larsson Syndrome enzyme fatty aldehyde dehydrogenase.
Keller, M.A., Zander, U., Fuchs, J.E., Kreutz, C., Watschinger, K., Mueller, T., Golderer, G., Liedl, K.R., Ralser, M., Krautler, B., Werner, E.R., Marquez, J.A.(2014) Nat Commun 5: 4439-4439
- PubMed: 25047030 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms5439
- Primary Citation Related Structures:
4QGK - PubMed Abstract:
Mutations in the gene coding for membrane-bound fatty aldehyde dehydrogenase (FALDH) lead to toxic accumulation of lipid species and development of the Sjögren-Larsson Syndrome (SLS), a rare disorder characterized by skin defects and mental retardation. Here, we present the crystallographic structure of human FALDH, the first model of a membrane-associated aldehyde dehydrogenase. The dimeric FALDH displays a previously unrecognized element in its C-terminal region, a 'gatekeeper' helix, which extends over the adjacent subunit, controlling the access to the substrate cavity and helping orientate both substrate cavities towards the membrane surface for efficient substrate transit between membranes and catalytic site. Activity assays demonstrate that the gatekeeper helix is important for directing the substrate specificity of FALDH towards long-chain fatty aldehydes. The gatekeeper feature is conserved across membrane-associated aldehyde dehydrogenases. Finally, we provide insight into the previously elusive molecular basis of SLS-causing mutations.
- 1] Division of Biological Chemistry, Biocenter, Innsbruck Medical University, Innrain 80-82, 6020 Innsbruck, Austria [2] Department of Biochemistry and Cambridge Systems Biology Centre, University of Cambridge, 80 Tennis court Rd, Cambridge CB2 1GA, UK.
Organizational Affiliation: