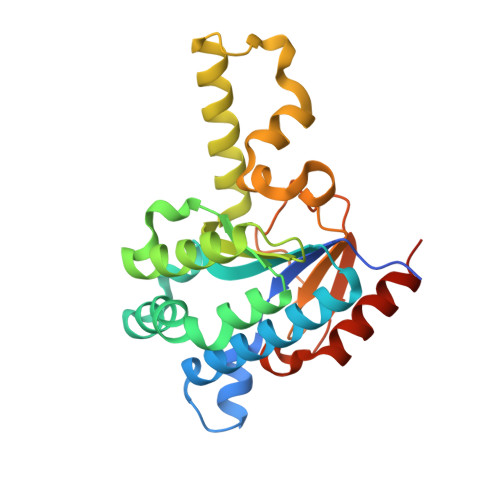
Discovery and structural characterization of an allosteric inhibitor of bacterial cis-prenyltransferase.
Danley, D.E., Baima, E.T., Mansour, M., Fennell, K.F., Chrunyk, B.A., Mueller, J.P., Liu, S., Qiu, X.(2015) Protein Sci 24: 20-26
- PubMed: 25287857 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2579
- Primary Citation Related Structures:
4Q9M, 4Q9O - PubMed Abstract:
Undecaprenyl pyrophosphate synthase (UPPs) is an essential enzyme in a key bacterial cell wall synthesis pathway. It catalyzes the consecutive condensations of isopentenyl pyrophosphate (IPP) groups on to a trans-farnesyl pyrophosphate (FPP) to produce a C55 isoprenoid, undecaprenyl pyrophosphate (UPP). Here we report the discovery and co-crystal structures of a drug-like UPPs inhibitor in complex with Streptococcus pneumoniae UPPs, with and without substrate FPP, at resolutions of 2.2 and 2.1 Å, respectively. The UPPs inhibitor has a low molecular weight (355 Da), but displays potent inhibition of UPP synthesis in vitro (IC50 50 nM) that translates into excellent whole cell antimicrobial activity against pathogenic strains of Streptococcal species (MIC90 0.4 µg mL(-1) ). Interestingly, the inhibitor does not compete with the substrates but rather binds at a site adjacent to the FPP binding site and interacts with the tail of the substrate. Based on the structures, an allosteric inhibition mechanism of UPPs is proposed for this inhibitor. This inhibition mechanism is supported by biochemical and biophysical experiments, and provides a basis for the development of novel antibiotics targeting Streptococcus pneumoniae.
- Worldwide Research and Development, Pfizer Inc, Groton, Connecticut, 06340.
Organizational Affiliation: