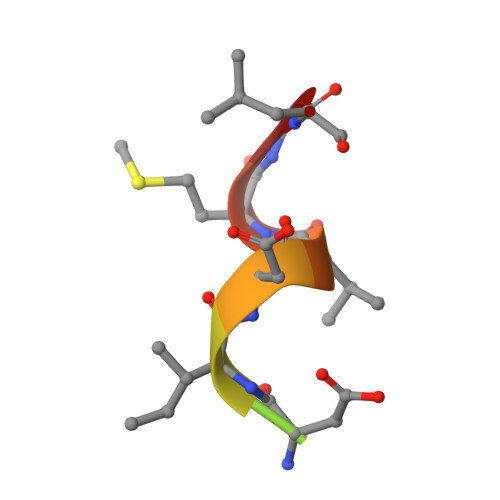
Crystal structure of a common GPCR-binding interface for G protein and arrestin.
Szczepek, M., Beyriere, F., Hofmann, K.P., Elgeti, M., Kazmin, R., Rose, A., Bartl, F.J., von Stetten, D., Heck, M., Sommer, M.E., Hildebrand, P.W., Scheerer, P.(2014) Nat Commun 5: 4801-4801
- PubMed: 25205354 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms5801
- Primary Citation Related Structures:
4PXF - PubMed Abstract:
G-protein-coupled receptors (GPCRs) transmit extracellular signals to activate intracellular heterotrimeric G proteins (Gαβγ) and arrestins. For G protein signalling, the Gα C-terminus (GαCT) binds to a cytoplasmic crevice of the receptor that opens upon activation. A consensus motif is shared among GαCT from the Gi/Gt family and the 'finger loop' region (ArrFL1-4) of all four arrestins. Here we present a 2.75 Å crystal structure of ArrFL-1, a peptide analogue of the finger loop of rod photoreceptor arrestin, in complex with the prototypical GPCR rhodopsin. Functional binding of ArrFL to the receptor was confirmed by ultraviolet-visible absorption spectroscopy, competitive binding assays and Fourier transform infrared spectroscopy. For both GαCT and ArrFL, binding to the receptor crevice induces a similar reverse turn structure, although significant structural differences are seen at the rim of the binding crevice. Our results reflect both the common receptor-binding interface and the divergent biological functions of G proteins and arrestins.
- Institut für Medizinische Physik und Biophysik (CC2), Charité-Universitätsmedizin Berlin, Charitéplatz 1, D-10117 Berlin, Germany.
Organizational Affiliation: