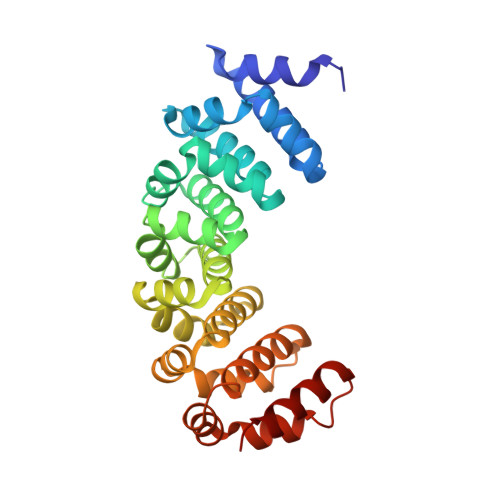
Crystal structures of designed armadillo repeat proteins: Implications of construct design and crystallization conditions on overall structure.
Reichen, C., Madhurantakam, C., Pluckthun, A., Mittl, P.R.(2014) Protein Sci 23: 1572-1583
- PubMed: 25132085
- DOI: https://doi.org/10.1002/pro.2535
- Primary Citation Related Structures:
4PLQ, 4PLR, 4PLS - PubMed Abstract:
Designed armadillo repeat proteins (dArmRP) are promising modular proteins for the engineering of binding molecules that recognize extended polypeptide chains. We determined the structure of a dArmRP containing five internal repeats and 3rd generation capping repeats in three different states by X-ray crystallography: without N-terminal His6 -tag and in the presence of calcium (YM5 A/Ca(2+) ), without N-terminal His6 -tag and in the absence of calcium (YM5 A), and with N-terminal His6 -tag and in the presence of calcium (His-YM5 A/Ca(2+)). All structures show different quaternary structures and superhelical parameters. His-YM5 A/Ca(2+) forms a crystallographic dimer, which is bridged by the His6 -tag, YM5 A/Ca(2+) forms a domain-swapped tetramer, and only in the absence of calcium and the His6 -tag, YM5 A forms a monomer. The changes of superhelical parameters are a consequence of calcium binding, because calcium ions interact with negatively charged residues, which can also participate in the modulation of helix dipole moments between adjacent repeats. These observations are important for further optimizations of dArmRPs and provide a general illustration of how construct design and crystallization conditions can influence the exact structure of the investigated protein.
- Department of Biochemistry, University of Zürich, Winterthurerstrasse 190, 8057, Zürich, Switzerland.
Organizational Affiliation: