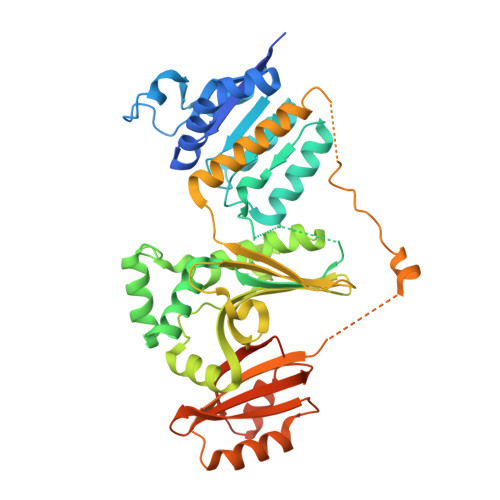
Structural basis for the catalytic mechanism of homoserine dehydrogenase.
Navratna, V., Reddy, G., Gopal, B.(2015) Acta Crystallogr D Biol Crystallogr 71: 1216-1225
- PubMed: 25945586 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004715004617
- Primary Citation Related Structures:
4PG4, 4PG5, 4PG6, 4PG7, 4PG8 - PubMed Abstract:
Homoserine dehydrogenase (HSD) is an oxidoreductase in the aspartic acid pathway. This enzyme coordinates a critical branch point of the metabolic pathway that leads to the synthesis of bacterial cell-wall components such as L-lysine and m-DAP in addition to other amino acids such as L-threonine, L-methionine and L-isoleucine. Here, a structural rationale for the hydride-transfer step in the reaction mechanism of HSD is reported. The structure of Staphylococcus aureus HSD was determined at different pH conditions to understand the basis for the enhanced enzymatic activity at basic pH. An analysis of the crystal structure revealed that Lys105, which is located at the interface of the catalytic and cofactor-binding sites, could mediate the hydride-transfer step of the reaction mechanism. The role of Lys105 was subsequently confirmed by mutational analysis. Put together, these studies reveal the role of conserved water molecules and a lysine residue in hydride transfer between the substrate and the cofactor.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore, Karnataka 560 012, India.
Organizational Affiliation: