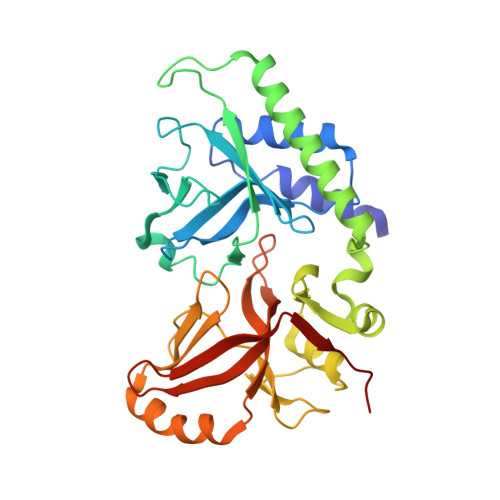
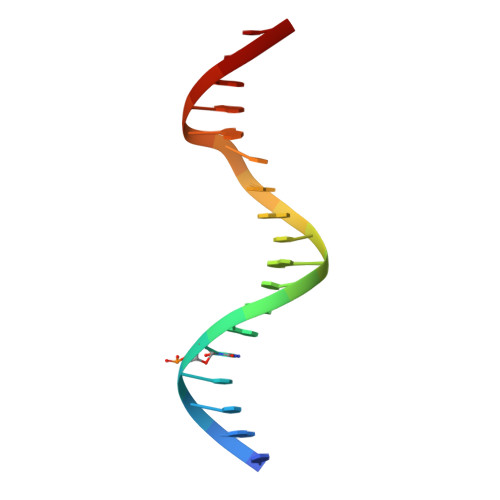
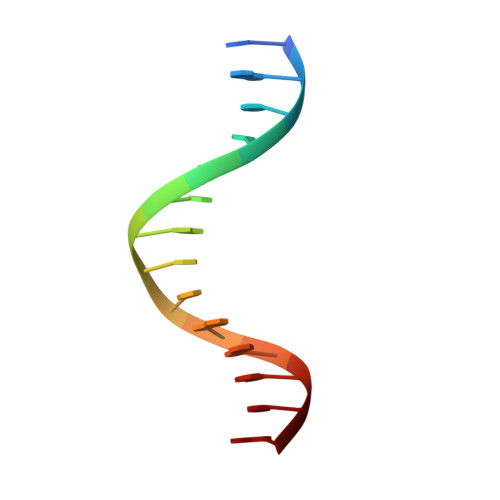
Structure of 5-hydroxymethylcytosine-specific restriction enzyme, AbaSI, in complex with DNA.
Horton, J.R., Borgaro, J.G., Griggs, R.M., Quimby, A., Guan, S., Zhang, X., Wilson, G.G., Zheng, Y., Zhu, Z., Cheng, X.(2014) Nucleic Acids Res 42: 7947-7959
- PubMed: 24895434
- DOI: https://doi.org/10.1093/nar/gku497
- Primary Citation Related Structures:
4PAR, 4PBA, 4PBB - PubMed Abstract:
AbaSI, a member of the PvuRts1I-family of modification-dependent restriction endonucleases, cleaves deoxyribonucleic acid (DNA) containing 5-hydroxymethylctosine (5hmC) and glucosylated 5hmC (g5hmC), but not DNA containing unmodified cytosine. AbaSI has been used as a tool for mapping the genomic locations of 5hmC, an important epigenetic modification in the DNA of higher organisms. Here we report the crystal structures of AbaSI in the presence and absence of DNA. These structures provide considerable, although incomplete, insight into how this enzyme acts. AbaSI appears to be mainly a homodimer in solution, but interacts with DNA in our structures as a homotetramer. Each AbaSI subunit comprises an N-terminal, Vsr-like, cleavage domain containing a single catalytic site, and a C-terminal, SRA-like, 5hmC-binding domain. Two N-terminal helices mediate most of the homodimer interface. Dimerization brings together the two catalytic sites required for double-strand cleavage, and separates the 5hmC binding-domains by ∼70 Å, consistent with the known activity of AbaSI which cleaves DNA optimally between symmetrically modified cytosines ∼22 bp apart. The eukaryotic SET and RING-associated (SRA) domains bind to DNA containing 5-methylcytosine (5mC) in the hemi-methylated CpG sequence. They make contacts in both the major and minor DNA grooves, and flip the modified cytosine out of the helix into a conserved binding pocket. In contrast, the SRA-like domain of AbaSI, which has no sequence specificity, contacts only the minor DNA groove, and in our current structures the 5hmC remains intra-helical. A conserved, binding pocket is nevertheless present in this domain, suitable for accommodating 5hmC and g5hmC. We consider it likely, therefore, that base-flipping is part of the recognition and cleavage mechanism of AbaSI, but that our structures represent an earlier, pre-flipped stage, prior to actual recognition.
- Department of Biochemistry, Emory University School of Medicine, 1510 Clifton Road, Atlanta, Georgia 30322, USA.
Organizational Affiliation: