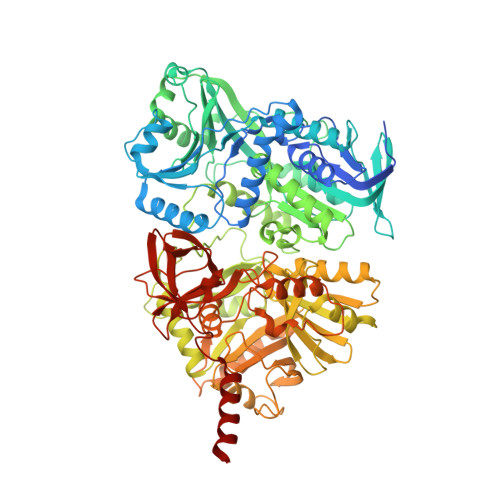
Folate in demethylation: The crystal structure of the rat dimethylglycine dehydrogenase complexed with tetrahydrofolate.
Luka, Z., Pakhomova, S., Loukachevitch, L.V., Newcomer, M.E., Wagner, C.(2014) Biochem Biophys Res Commun 449: 392-398
- PubMed: 24858690 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bbrc.2014.05.064
- Primary Citation Related Structures:
4P9S, 4PAA, 4PAB - PubMed Abstract:
Dimethylglycine dehydrogenase (DMGDH) is a mammalian mitochondrial enzyme which plays an important role in the utilization of methyl groups derived from choline. DMGDH is a flavin containing enzyme which catalyzes the oxidative demethylation of dimethylglycine in vitro with the formation of sarcosine (N-methylglycine), hydrogen peroxide and formaldehyde. DMGDH binds tetrahydrofolate (THF) in vivo, which serves as an acceptor of formaldehyde and in the cell the product of the reaction is 5,10-methylenetetrahydrofolate instead of formaldehyde. To gain insight into the mechanism of the reaction we solved the crystal structures of the recombinant mature and precursor forms of rat DMGDH and DMGDH-THF complexes. Both forms of DMGDH reveal similar kinetic parameters and have the same tertiary structure fold with two domains formed by N- and C-terminal halves of the protein. The active center is located in the N-terminal domain while the THF binding site is located in the C-terminal domain about 40Å from the isoalloxazine ring of FAD. The folate binding site is connected with the enzyme active center via an intramolecular channel. This suggests the possible transfer of the intermediate imine of dimethylglycine from the active center to the bound THF where they could react producing a 5,10-methylenetetrahydrofolate. Based on the homology of the rat and human DMGDH the structural basis for the mechanism of inactivation of the human DMGDH by naturally occurring His109Arg mutation is proposed.
- Department of Biochemistry, Vanderbilt University Medical Center, Nashville, TN 37232, USA. Electronic address: z.luka@vanderbilt.edu.
Organizational Affiliation: