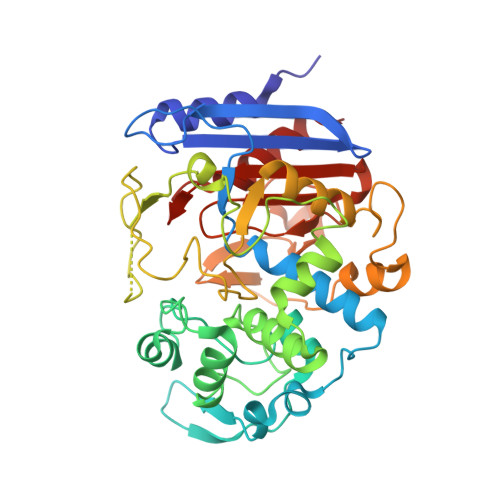
Crystallographic analysis and biochemical applications of a novel penicillin-binding protein/ beta-lactamase homologue from a metagenomic library.
Ngo, T.D., Ryu, B.H., Ju, H., Jang, E.J., Kim, K.K., Kim, T.D.(2014) Acta Crystallogr D Biol Crystallogr 70: 2455-2466
- PubMed: 25195758
- DOI: https://doi.org/10.1107/S1399004714015272
- Primary Citation Related Structures:
4P6B, 4P85, 4P87 - PubMed Abstract:
Interest in penicillin-binding proteins and β-lactamases (the PBP-βL family) is increasing owing to their biological and clinical significance. In this study, the crystal structure of Est-Y29, a metagenomic homologue of the PBP-βL family, was determined at 1.7 Å resolution. In addition, complex structures of Est-Y29 with 4-nitrophenyl phosphate (4NP) and with diethyl phosphonate (DEP) at 2.0 Å resolution were also elucidated. Structural analyses showed that Est-Y29 is composed of two domains: a β-lactamase fold and an insertion domain. A deep hydrophobic patch between these domains defines a wide active site, and a nucleophilic serine (Ser58) residue is located in a groove defined primarily by hydrophobic residues between the two domains. In addition, three hydrophobic motifs, which make up the substrate-binding site, allow this enzyme to hydrolyze a wide variety of hydrophobic compounds, including fish and olive oils. Furthermore, cross-linked Est-Y29 aggregates (CLEA-Est-Y29) significantly increase the stability of the enzyme as well as its potential for extensive reuse in various deactivating conditions. The structural features of Est-Y29, together with biochemical and biophysical studies, could provide a molecular basis for understanding the properties and regulatory mechanisms of the PBP-βL family and their potential for use in industrial biocatalysts.
- Department of Molecular Cell Biology, Samsung Biomedical Research Institute, Sungkyunkwan University School of Medicine, Suwon 440-746, Republic of Korea.
Organizational Affiliation: