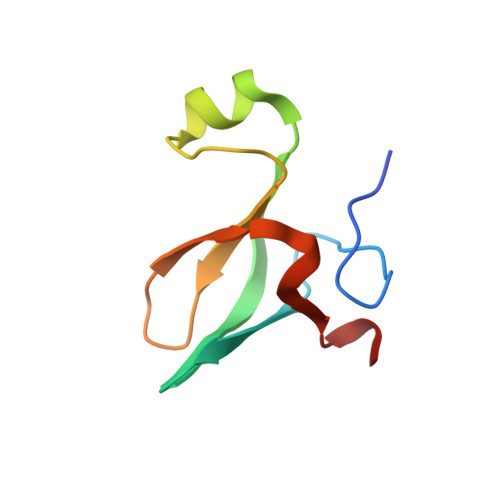
A Small Protein Associated with Fungal Energy Metabolism Affects the Virulence of Cryptococcus neoformans in Mammals.
McClelland, E.E., Ramagopal, U.A., Rivera, J., Cox, J., Nakouzi, A., Prabu, M.M., Almo, S.C., Casadevall, A.(2016) PLoS Pathog 12: e1005849-e1005849
- PubMed: 27583447
- DOI: https://doi.org/10.1371/journal.ppat.1005849
- Primary Citation Related Structures:
4P5N - PubMed Abstract:
The pathogenic yeast Cryptococcus neoformans causes cryptococcosis, a life-threatening fungal disease. C. neoformans has multiple virulence mechanisms that are non-host specific, induce damage and interfere with immune clearance. Microarray analysis of C. neoformans strains serially passaged in mice associated a small gene (CNAG_02591) with virulence. This gene, hereafter identified as HVA1 (hypervirulence-associated protein 1), encodes a protein that has homologs of unknown function in plant and animal fungi, consistent with a conserved mechanism. Expression of HVA1 was negatively correlated with virulence and was reduced in vitro and in vivo in both mouse- and Galleria-passaged strains of C. neoformans. Phenotypic analysis in hva1Δ and hva1Δ+HVA1 strains revealed no significant differences in established virulence factors. Mice infected intravenously with the hva1Δ strain had higher fungal burden in the spleen and brain, but lower fungal burden in the lungs, and died faster than mice infected with H99W or the hva1Δ+HVA1 strain. Metabolomics analysis demonstrated a general increase in all amino acids measured in the disrupted strain and a block in the TCA cycle at isocitrate dehydrogenase, possibly due to alterations in the nicotinamide cofactor pool. Macrophage fungal burden experiments recapitulated the mouse hypervirulent phenotype of the hva1Δ strain only in the presence of exogenous NADPH. The crystal structure of the Hva1 protein was solved, and a comparison of structurally similar proteins correlated with the metabolomics data and potential interactions with NADPH. We report a new gene that modulates virulence through a mechanism associated with changes in fungal metabolism.
- Department of Biology, Middle Tennessee State University, Murfreesboro, Tennessee, United States of America.
Organizational Affiliation: