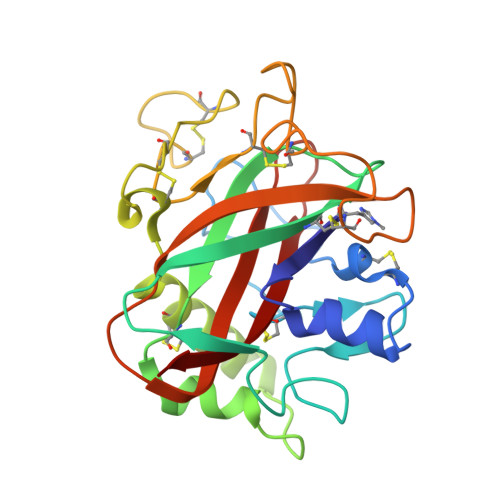
Structure and boosting activity of a starch-degrading lytic polysaccharide monooxygenase.
Lo Leggio, L., Simmons, T.J., Poulsen, J.C., Frandsen, K.E., Hemsworth, G.R., Stringer, M.A., von Freiesleben, P., Tovborg, M., Johansen, K.S., De Maria, L., Harris, P.V., Soong, C.L., Dupree, P., Tryfona, T., Lenfant, N., Henrissat, B., Davies, G.J., Walton, P.H.(2015) Nat Commun 6: 5961-5961
- PubMed: 25608804 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms6961
- Primary Citation Related Structures:
4OPB - PubMed Abstract:
Lytic polysaccharide monooxygenases (LPMOs) are recently discovered enzymes that oxidatively deconstruct polysaccharides. LPMOs are fundamental in the effective utilization of these substrates by bacteria and fungi; moreover, the enzymes have significant industrial importance. We report here the activity, spectroscopy and three-dimensional structure of a starch-active LPMO, a representative of the new CAZy AA13 family. We demonstrate that these enzymes generate aldonic acid-terminated malto-oligosaccharides from retrograded starch and boost significantly the conversion of this recalcitrant substrate to maltose by β-amylase. The detailed structure of the enzyme's active site yields insights into the mechanism of action of this important class of enzymes.
- Department of Chemistry, University of Copenhagen, Universitetsparken 5, 2100 Copenhagen Ø, Denmark.
Organizational Affiliation: